Comparison of support vector machines based on particle swarm optimization and genetic algorithm in sleep staging
Abstract
BACKGROUND:
Heart rate variability (HRV) can reflect the relationship between heart rhythm and sleep structure.
OBJECTIVE:
In order to study the effect of support vector machine (SVM) on the results of automatic sleep staging and improve the effectiveness of heart rate variability (HRV) as a sleep structure biomarker, thereby realize long term and non-contact monitoring of sleep quality.
METHODS:
Two kinds of parameter optimization methods are applied to stage sleep experiments when the known SVM can be used for automatic sleep staging. By factor analysis of the time domain, frequency domain, and nonlinear dynamic characteristics of subjects’ HRV signals, the accuracy of the cross-validation method (K-CV) is used as the fitness function value in genetic algorithm (GA) and particle swarm optimization (PSO). Furthermore, GA and PSO are used to optimize the SVM parameters.
RESULTS:
The results show that the accuracy rate of sleep stage is 64.44% when parameters are not optimized, the accuracy rate based on PSO is improved to 78.89% and the accuracy rate based on GA is improved to 84.44%.
CONCLUSION:
Both optimization algorithms can improve the accuracy of SVM for sleep staging and better results based on GA in the experiment.
1.Introduction
Sleep quality affects human health, while sleep staging can assess sleep quality and related diseases. At present, the traditional sleep staging is based on electroencephalogram (EEG) as the golden standard for judging sleep. Oropesa and Cycon [1] use wavelet packet transformation (WPT) to sample EEG records for a long time of 30 seconds, and a feature generator to quantize and reduce the size of data sets, and an artificial neural network for optimal classification. Compared with human experts, the classification results reached a consistency of 70% to 80%. Liu et al. [2] proposed a novel method based on fuzzy entropy (FE) for the staging of EEG sleep characteristics and experimental results demonstrate that average identification rate reaches 87.1%. However, such methods require long contact between electrodes and the human body, which can easily affect natural sleep. Therefore, it is a hot research topic to find a method of automatic sleep staging for non EEG signals.
HRV refers to the minute change of the cardiac cycle, mainly affected by sympathetic and parasympathetic nerves. The time between two R wave in the successive heartbeats is called the RR interval (RR). It has been experimentally shown that HRV has an important relationship with sleep staging and physical health [3, 4, 5]. Versace et al. [6] makes global research on HRV at each stage and then uses wavelet transform to isolate very low-frequency (VLF), Low-frequency (LF) and High-frequency (HF) components, and estimates the nonlinear dynamics of HRV with sample entropy (SampEn). Wang et al. [7] used the support vector machine (SVM) to achieve good results in sleep stage classification by using principal component analysis (PCA) to reduce the dimensionality of multi-feature indicators. Liu et al. [8] extracted three parameters including symbol entropy index, detrended volatility index, and delta band energy ratio, combined with least squares support vector machine classifier to perform sample training fitting and classification recognition, and the accuracy of sleep staging can reach 92.87%. Due to the good performance of SVM in sleep stages, it is necessary to explore the influence of SVM parameters on sleep staging in order to make SVM more widely available. In this experiment, particle swarm optimization (PSO) and genetic algorithm (GA) were used to optimize SVM parameters [9, 10, 11]. Firstly, the HRV analysis of the electrocardiogram (ECG) signal was performed to extract the time-frequency and nonlinear dynamic characteristics of the data. Secondly, the characteristics were then reduced according to the result of the factor analysis. Finally, the optimal parameters of SVM were used to stage the sleep and the staging results are observed.
2.Materials
2.1Support vector machine (SVM)
SVM is a kind of theory and risk structure minimization and its learning method evolved from the linear separability problem. The SVM method based on statistical learning theory has a great advantage in pattern classification, and has been successfully applied to fault diagnosis, and the algorithm itself has been improved continuously [12, 13, 14, 15]. A rapid and effective SVM pattern recognition and regression software package LIBSVM developed by Professor Lin Chih-Jen of Taiwan University used in this experiment. The classifier model is an one-to-one (One-versus-one) method, in which the idea is to design an SVM between any two samples, that is, K (k-1)/2 SVM should be designed. When classifying an unknown sample, the category with the most votes is the class of the unknown sample. The training model of SVM is as follows.
1) Known training set:
where,
2) Select appropriate kernel function and appropriate parameters to construct and solve the optimization problem:
Get the best solution:
3) Calculate thresholds
Where,
4) Construct decision function
2.2Particle swarm optimization (PSO)
PSO is an optimization algorithm based on swarm intelligence, which is an ant colony algorithm in the field of computer intelligence. Based on the stochastic solution and the iterative search for the optimal solution, the PSO also evaluates the quality of the solution through the adaptability. But it is simpler than the rule of genetic algorithm, it has no Crossover and Mutation operation of Genetic algorithm. It finds the global optimal by following the best solution that is currently searched. This kind of algorithm has attracted much attention from the academic community because of its advantages of easy implementation, high precision and fast convergence. It is a parallel algorithm.
The principle of PSO is as follows.
where,
2.3Genetic algorithm (GA)
GA originated from the computer simulation of biological systems. Professor Holland and his students in the United States are inspired by biological simulation techniques to create an adaptive probabilistic optimization technique for complex system optimization based on biological genetics and evolutionary mechanism. It introduces the biological evolution principle of nature into the system of optimizing parameters coding optimizing parameters, according to the selected fitness function and through genetic selection, crossover and mutation of the individual screening, so that the fitness value of the individual is retained and the adaptation of the poor individual is eliminated. The new group inherits the information of the previous generation and is superior to the previous generation. The basic operation of genetic algorithm is as follows:
1. Select operation: It refers to the probability and fitness value of the individual selected from the old group to the new group with certain probability. The better the adaptability of the individual, the greater the probability of being chosen.
2. Crossover operation: It refers to the selection of two individuals from the individual and the exchange of two chromosomes to produce new outstanding individuals. The crossover process is to exchange two chromosomes from a group, randomly select a point or point chromosome position.
3. Mutation operation: Mutation operation is to choose an individual from the group, then select a point in the chromosome to mutate to produce very good individuals.
2.4Algorithm for parameter adjustment of SVM
When using SVM to do classified prediction, it is necessary to adjust the relevant parameters to get the ideal accuracy of prediction classification. The idea of using CV can obtain optimal parameters in a certain sense, which can effectively avoid the occurrence of over-learning and under-learning. Finally, the prediction accuracy of the test set is better. The experiments show that the model chosen by CV to train SVM is more effective than the random selection of parameter training SVM. The K-CV Cross validation method is selected in this experiment. That is, the accuracy of the training set in the sense of CV is used as the fitness function value in PSO and GA. the whole algorithm of optimizing SVM parameters by PSO and GA as show in Figs 1 and 2.
Figure 1.
Algorithm flowchart for optimizing SVM parameters using PSO.
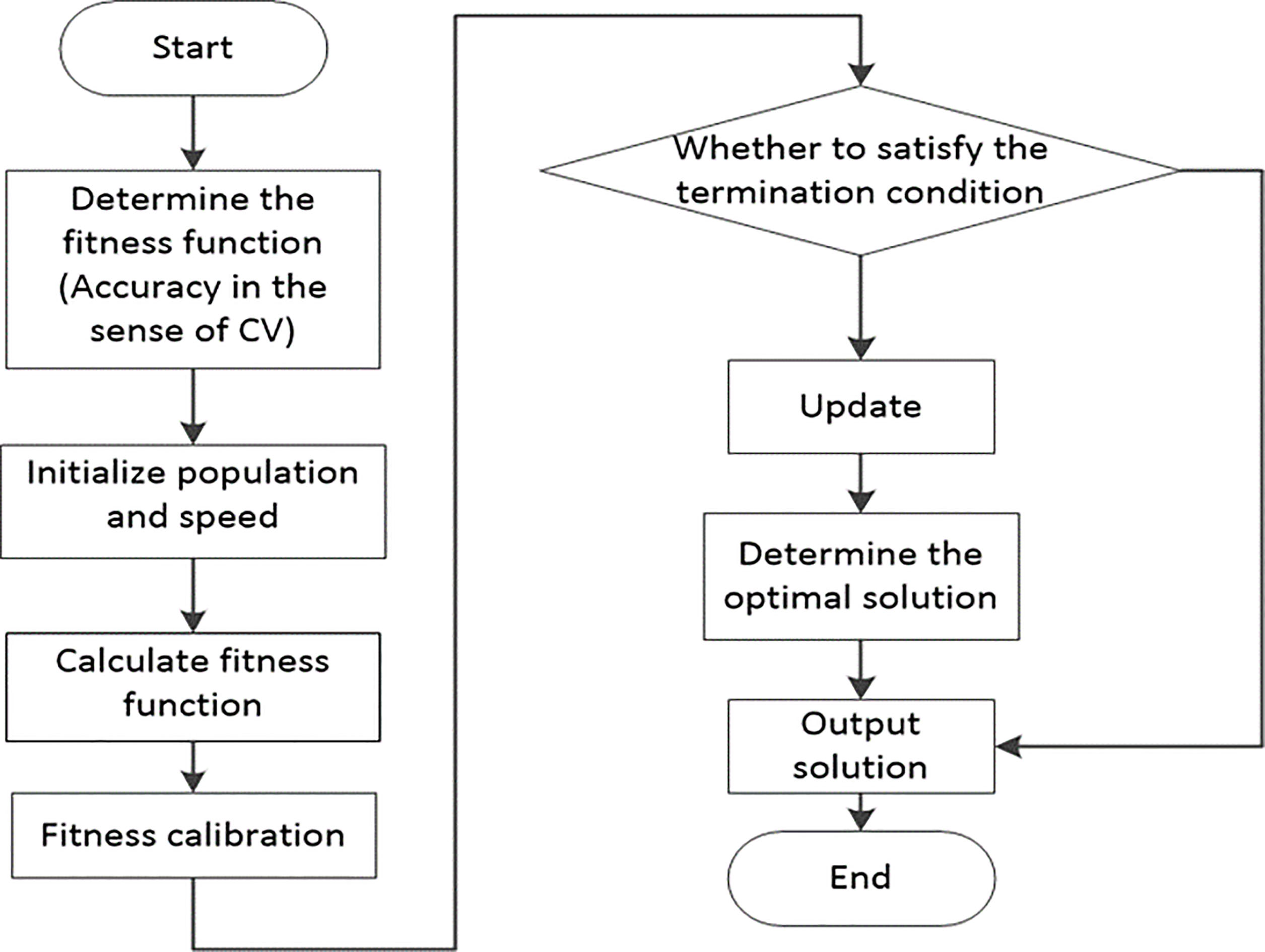
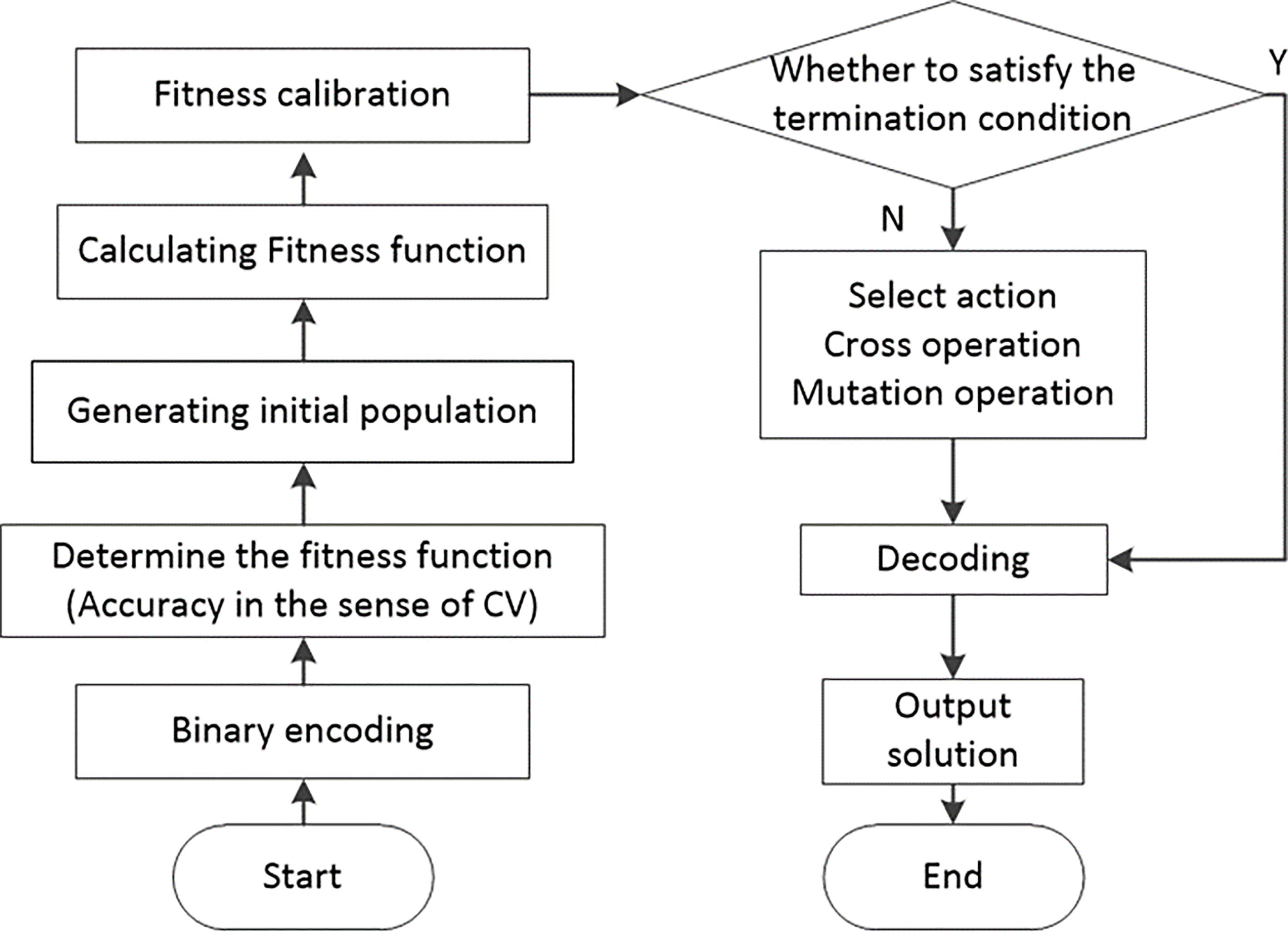
Figure 2.
Algorithm flowchart for optimizing SVM parameters using GA.
3.Methods
The experimental data is the original ECG signal in the MIT-BIH polysomnography database. The sampling frequency is 250 HZ. All ECG signals are divided into segments for each 30 s and are staged by experts. ECG signals were extracted from five individuals for HRV analysis. Samples were analyzed according to wake (W), non-rapid eye movement (NREM), and rapid eye movement (REM) groups. The sample size was 120. Since there have been literatures confirming that heart rate changes occur before the sleep stage, in order to be able to effectively observe the effect of SVM parameters on sleep stage outcomes, continuous multiple stages are selected to be the same stage of data [16, 17, 18]. Then perform factor analysis on the results to reduce the data redundancy. Replace the original characteristic quantity with the principal component with a cumulative contribution rate exceeding 93.799%.
Three groups of control experiments were designed to verify the effect of parameter optimization on sleep staging. Experiment one: Using SVM directly to stage the sleep. Experiment two: The sleep staging of SVM based on PSO parameters optimization. Experiment three: The sleep staging of SVM based on GA parameters optimization. In this experiment, the kernel function used by SVM is radial basis function (RBF).
4.Results
4.1Pre-processing of RR intervals
Preprocessing is to produce a sequence of observations from the RR interval sequence (RIS), and then to analyze it by HRV. We first remove some anomalous RR interval values (RR interval
Table 1
Total variance explained
| Component | Total | Variance/% | Cumulative/% |
|---|---|---|---|
| 1 | 4.640 | 58.003 | 58.003 |
| 2 | 1.795 | 21.986 | 79.989 |
| 3 | 1.105 | 13.810 | 93.799 |
| 4 | 0.368 | 4.605 | 98.403 |
| 5 | 0.085 | 1.064 | 99.467 |
| 6 | 0.043 | 0.533 | 100.000 |
| 7 | 0.000 | 0.000 | 100.000 |
Figure 3.
Pre-processing of RR intervals.
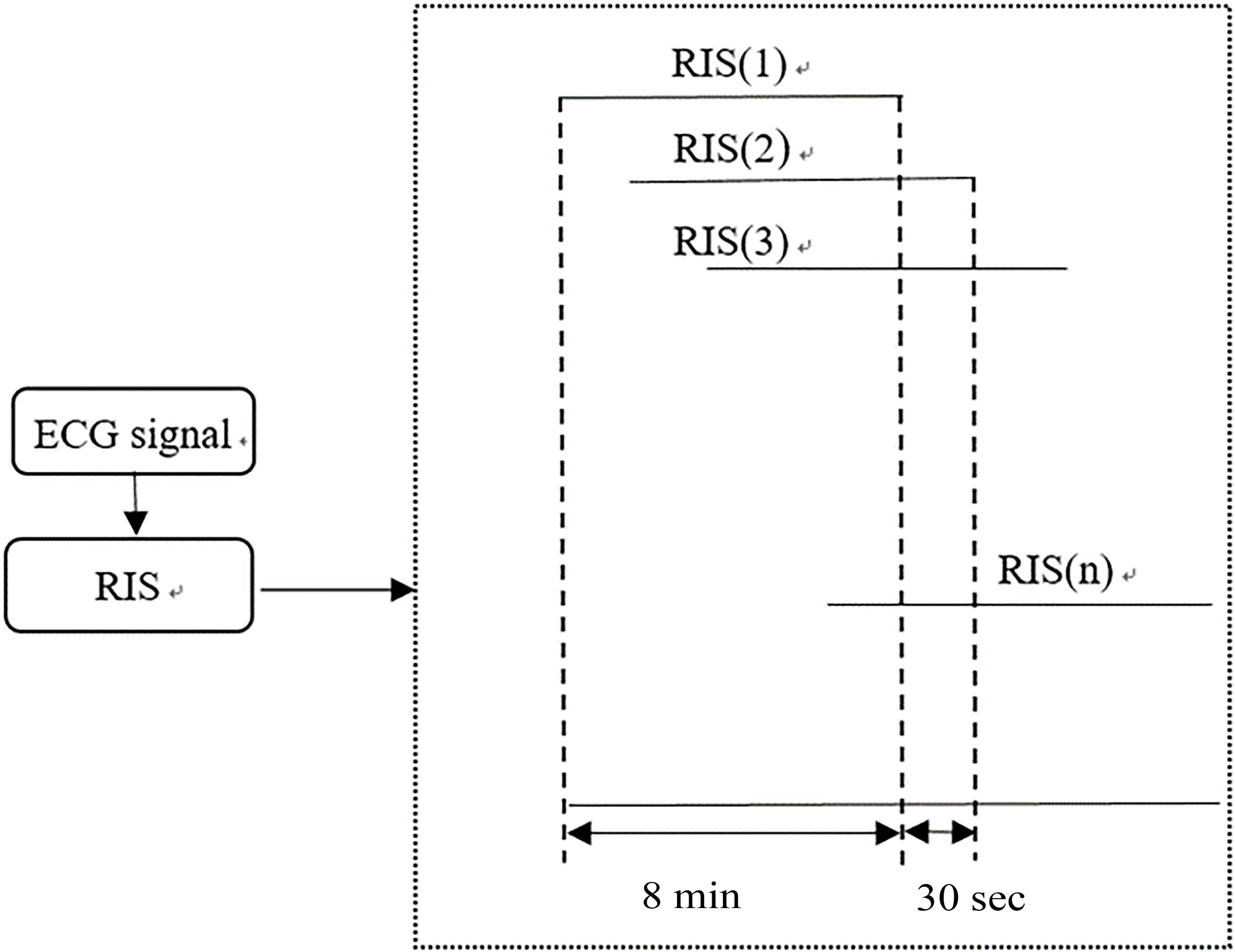
Figure 4.
Using SVM directly to stage the sleep.
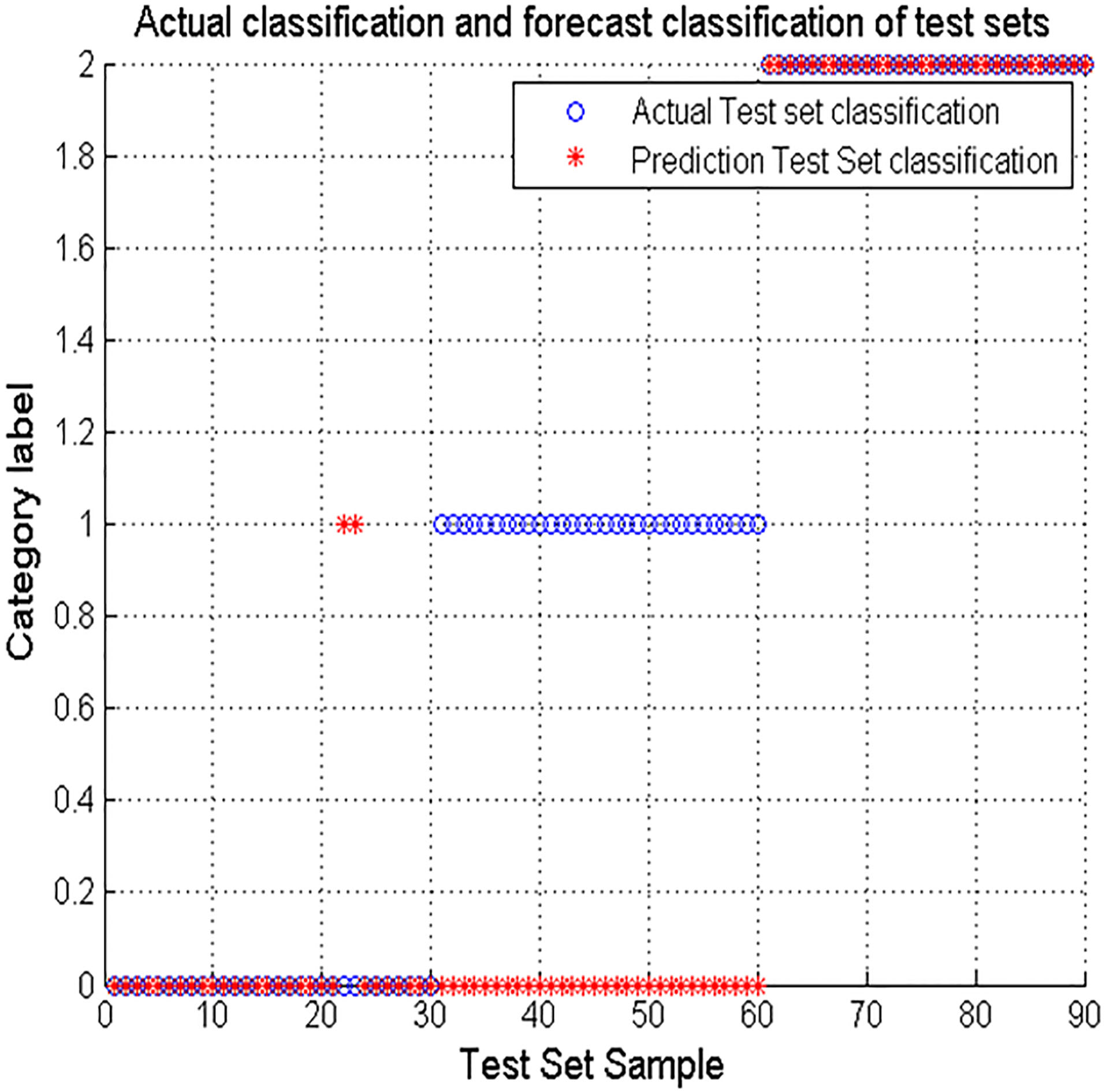
Figure 5.
Sleep stage results optimized based on PSO parameters.
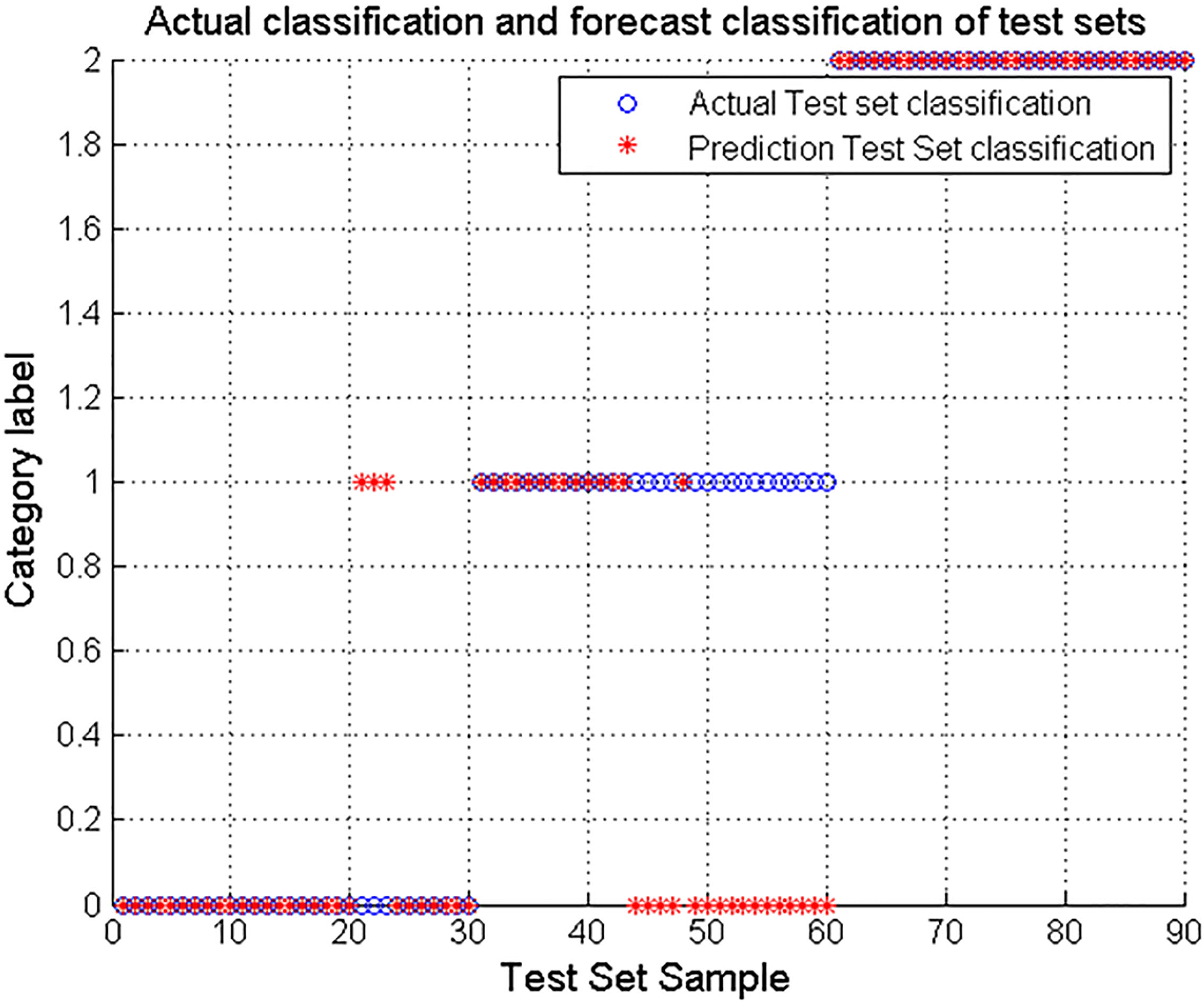
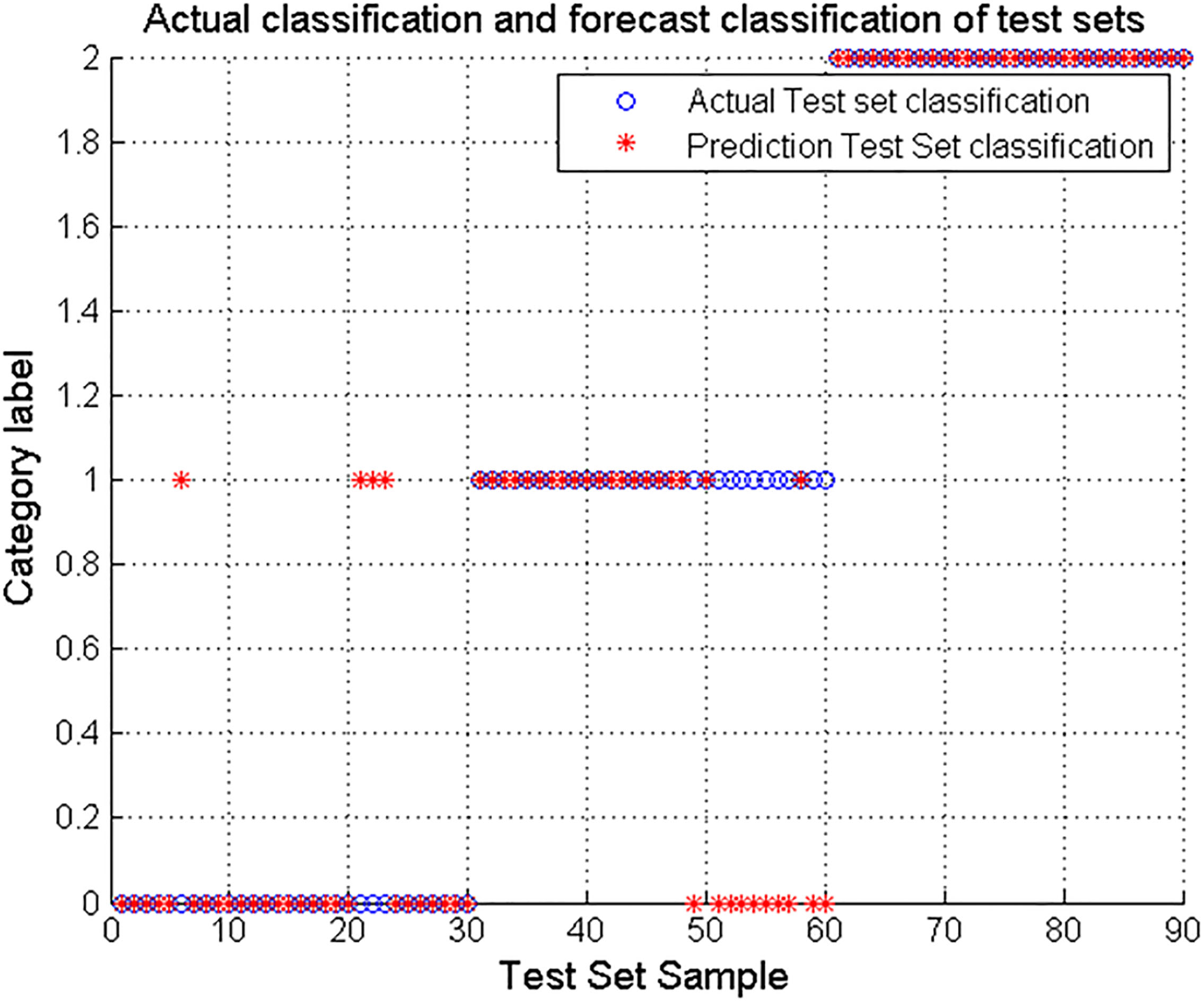
Figure 6.
Sleep stage results optimized based on GA parameters.
4.2The result of reducing dimension by factor analysis of HRV
The characteristics of HRV from the time domain, frequency domain and nonlinear dynamics were analyzed during the experiment. Then the standard deviation (SDNN) of the difference between the heart rate (HR), the average RR interval, all adjacent normal RR intervals, the root mean square of the difference between the normal adjacent RR intervals (RMSSD), PNN50, and low frequency were extracted. After the factor analysis, the cumulative contribution rate of each component as shown in Table 1.
4.3SVM parameter experiment results
The results show that ‘o’ represents the actual test set classification, ‘*’ Indicates the prediction test set classification. ‘0’ indicates the Wake period, ‘1’ means Nrem period and ‘2’ indicates REM period. As can be seen from the data in Fig. 4, when the SVM is not optimized for the parameters, the accuracy is the lowest, only 64.44%. In the three-period results, the results of NREM were the worst, almost 0. Figure 5 shows that after PSO parameter optimization, the accuracy of SVM for sleep staging has increased to 78.89%. The classification result for the Wake period is the worst, only 46.67%. Figure 6 shows that after the optimization of the GA parameters, the accuracy of SVM for sleep staging is improved to 84.44%. Among them, the accuracy rate of Wake and NREM is 86.67% and 66.67% respectively. In summary, the results show that the parameter-optimized SVM improves the accuracy of sleep staging, and this experiment shows that the optimization based on GA parameters is better.
5.Discussion
This experiment demonstrated the effect of SVM parameters on the accuracy of sleep staging and concluded that SVM with optimized GA parameters had better effect on sleep staging. However, GA is more complex and PSO is relatively simple. PSO may fall into a local optimal solution and GA can find a global optimum. Therefore, GA and PSO can be combined in future explorations to avoid their respective disadvantages [19, 20]. The purpose of the experiment is to further promote the application of SVM in sleep staging, the results of classification should not only consider the accuracy, but also consider the impact of individual differences on data processing. What’s more, because of the influence of the root SVM, the accuracy rate will decrease when the sleep stage increases, so this experiment only performs three stages of sleep. Therefore, it is necessary to further optimize the algorithm in subsequent experiments, so as to improve the accuracy of the five-stage sleep staging. Therefore, it is necessary to add more features such as body motion signals, respiratory signals, and oxygen saturation to improve the accuracy of the five-stage sleep staging.
Acknowledgments
The authors would like to thank Physionet for providing the Sleep-EDF database.
Conflict of interest
The authors declare that there is no conflict of interest regarding the publication of this article.
References
[1] | Oropesa E, Cycon HL. Sleep Stage Classification using Wavelet Transform and Neural Network. (1999) . |
[2] | Liu H, Xie H, He W. Characterization and classification of eeg sleep stage based on fuzzy entropy. Journal of Data Acquisition and Processing, (2010) , 25: (4): 484-489. |
[3] | Burns JW, Crofford LJ, Chervin RD. Sleep stage dynamics in fibromyalgia patients and controls. Sleep Medicine, (2007) , 9: (6). |
[4] | Kishi A, Natelson BH, Togo F, et al. Sleep-stage dynamics in patients with chronic fatigue syndrome with or without fibromyalgia. Sleep, (2011) , 34: (11): 1551. |
[5] | Bruck MLJD. Sleep abnormalities in chronic fatigue syndrome/myalgic encephalomyelitis: a review. Journal of Clinical Sleep Medicine Jcsm Official Publication of the American Academy of Sleep Medicine, (2012) , 8: (6): 719-28. |
[6] | Versace F, Mozzato M, De MTG, et al. Heart rate variability during sleep as a function of the sleep cycle. Biological Psychology, (2003) , 63: (2): 149-162. |
[7] | Wang J, Sun W, Wei R, et al. Study on sleep staging methods based on heart rate variability analysis. Journal of Biomedical Engineering, (2016) , 33: (3): 420-425. |
[8] | Liu Z, Zhang H, Zhao H, et al. Study on sleep staging algorithm based on EEG signals. Chinese Journal of Biomedical Engineering, (2015) , 34: (6): 693-700. |
[9] | Wang D, Lu C, Jiang W, et al. Study on PSO-based decision-tree SVM multi-class classification method. Journal of Electronic Measurement and Instrumentation, (2015) , 29: (4): 611-615. |
[10] | Chapelle O, Vapnik V, Bousquet O, et al. Choosing multiple parameters for support vector machines. Machine Learning, (2002) , 46: (1-3): 131-159. |
[11] | Huang HP, Liu YH. Fuzzy support vector machines for pattern recognition and data mining. Intl. J. of Fuzzy Systems, (2002) , 4: (3): 826-835. |
[12] | Jiao L, Bo L, Wang L. Fast sparse approximation for least squares support vector machine. IEEE Trans Neural Netw, (2007) , 18: (3): 685-97. |
[13] | Cui W, Pan J, He G, et al. Class center and feature weighting based feature selection algorithm. Electronic Measurement Technology, (2015) , 38: (3): 26-29. |
[14] | Yan B, Wang X, Jiang Z. Study on the gaze estimation algorithm based on support vector regression model. Chinese Journal of Scientific Instrument, (2014) , 35: (10): 2299-2305. |
[15] | Xia X-L, Jiao W, Li K, Irwin G, Zhang H. A novel sparse least squares support vector machines. Mathematical Problems in Engineering, (2013) , 2013. |
[16] | Otzenberger H, Simon C, Gronfier C, et al. Temporal relationship between dynamic heart rate variability and electroencephalographic activity during sleep in man. Neuroscience Letters, (1997) , 229: (3): 173-176. |
[17] | Zhuang Z, Gao S, Gao X. The sleep staging based on HRV analysis. Journal of Biomedical Engineering, (2006) , 23: (3): 499-504. |
[18] | Brandenberger G, Ehrhart J, Piquard F, et al. Inverse coupling between ultradian oscillations in delta wave activity and heart rate variability during sleep. Clinical Neurophysiology, (2001) , 112: (6): 992. |
[19] | Tsai CW, Tseng SP, Chiang MC, et al. A high-performance genetic algorithm: using traveling salesman problem as a case. Scientific World Journal, (2014) , 2014: (2): 178621. |
[20] | Shi XH, Wan LM, Lee HP, et al. An improved genetic algorithm with variable population-size and a PSO-GA based hybrid evolutionary algorithm// in: International Conference on Machine Learning and Cybernetics. IEEE, Vol. 3, (2003) : pp. 1735-1740. |




