PBRM1 Mutations as a Predictive Biomarker for Immunotherapy in Metastatic Renal Cell Carcinoma: A Systematic Review
Abstract
INTRODUCTION:
Genomic features linked to prediction of response to immunotherapy in metastatic renal cell carcinoma (mRCC) are still lacking. Protein polybromo-1 (PBRM1) mutations have been studied as a potential biomarker of clinical benefit, with conflicting published data so far.
MATERIAL AND METHODS:
This systematic review was guided by the standards of the PRISMA statement to identify studies involving mRCC, immunotherapy and mutations in PBRM1. The main objective was to assess the relationship between PBRM1 mutations and response to immune checkpoint inhibitors (ICI) in patients with mRCC.
RESULTS:
After an initial search that identified 422 studies, 8 studies met the eligibility criteria and were selected for the final analysis. Data are included from 2 trials in the first-line treatment setting, and 6 trials in second- or later treatment lines evaluating the relationship between the presence of PBRM1 mutations and clinical benefit (CB) with ICI treatment. Regarding the first-line treatment setting, the analysis of both studies failed to show any CB in patients with PBRM1 mutations treated with ICI. However, for the second- and later treatment lines, the results were mixed.
CONCLUSIONS:
PBRM1 mutations may be a potential genomic biomarker to predict response to ICI treatment in patients with mRCC, mainly in second- and later treatment lines, but the existence of conflicting data in the literature highlights an important bias in the studies and the need for additional clinical validation in large, prospective trials.
INTRODUCTION
Recent advances in the field of immuno-oncology (IO) have led the US Food and Drug Administration (FDA) to approve therapies based on immune checkpoint inhibitors (ICI) for different types of cancer, including metastatic renal cell carcinoma (mRCC) [1]. To date, however, immunotherapy has been shown to provide lasting clinical benefit (CB) to only a portion of patients. The identification of reliable biomarkers that could be a tool to select the patients who can most benefit from this systemic therapy is an unmet need.
The first biomarker identified as a predictor of good response to treatment with ICI in some types of neoplasms like lung, breast and gastric cancers was the expression of PD-L1, assessed by immunohistochemistry (IHC) [2]. More recently, other biomarkers such as high microsatellite instability (MSI-H) or mismatch repair deficiency (dMMR) and high tumor mutational burden (TMB) (defined as tumor mutation load > 10 mutations/megabase or TMB > 10 mut/Mb) have also shown association with superior efficacy and CB, leading the FDA to approve ICI as a tumor agnostic therapy [3, 4].
Unlike other types of tumors that respond to ICI, such as non-small cell lung cancer (NSCLC), melanoma and MSI-H colorectal adenocarcinoma, clear cell RCC (ccRCC) has a low TMB [5– 8]. While melanoma and NSCLC typically have 10 to 400 mut/Mb (1– 4), mRCC has an average of 1.1 mut/Mb [9, 10]. Despite this, ICI have been shown to prolong survival in a subset of patients with mRCC, leading to the hypothesis that different molecular mechanisms might be capable of making the tumor microenvironment (TME) immunologically active and therefore sensitive to ICI [11– 14]. However, so far, it is not clear what specific genomic characteristics of mRCC may be associated with this mechanism.
Mammalian switch/sucrose non-fermentable (SWI/SNF) complex is a key chromatin remodeler that regulates essential cellular processes such as transcription, DNA repair and replication. Mutations in SWI/SNF subunits have been described in several malignant tumors, but their impact on carcinogenesis and their potential therapeutic role are still unknown [15].
PBRM1 is one of the genes that compose PBAF (polybromo BRG1-associated factor), a subcomplex of the SWI/SNF family. Mutations in PBRM1 are found in about 7.7% of all malignant tumors, but in patients with ccRCC, these mutations can be present in up to about 46.6% of cases. PBRM1 mutations are more expressed in highly mutated tumors and approximately half of them (52.7%) are classified as loss-of-function (LOF) mutations, variants predicted to severely disrupt protein-coding genes. Truncating mutations like frameshift indels (insertion/deletion), nonsense (stop codon-introducing), and splice site-disrupting single nucleotide variants (SNVs) are the most likely to lead to loss of function of the protein [16].
PBRM1 expression is inversely correlated with T-cell cytotoxicity genes expression and may represent a mechanism of tumor cell resistance to ICI [16]. In animals and human models, previous data regarding the association between PBRM1 mutation and superior outcomes with ICI blockade are controversial. While in some trials PBRM1 LOF has been correlated with a more immunosuppressive and ICI-resistant microenvironment, in some animal models it changed the global expression profiles of tumor cells, increasing their sensitivity to ICI [17]. The mechanisms by which PBRM1 can interfere with the effectiveness of ICI are dependent of interferon responsive genes, JAK/STAT signaling and p53’transcriptional activity [18– 20].
In this context, we systematically review the current evidence on PBRM1 mutations as a potential predictive biomarker to ICI in patients with mRCC, and provide data on 8 clinical trials.
MATERIAL AND METHODS
Study design
The reporting of this systematic review was guided by the standards of the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) statement [21].
Search strategy
A systematic search was performed in 5 medical databases: 1) PubMed/MEDLINE, 2) Embase, 3) LILACS, 4) Cochrane, and 5) Scopus. The primary search included the keywords “renal cell carcinoma” AND “immunotherapy” AND “PBRM1”. The search was completed on 12 April 2021. A supplemental search was performed manually to identify published abstracts from the American Society of Clinical Oncology (ASCO) annual meetings, ASCO genito-urinary cancers symposiums and European Society for Medical Oncology annual meetings, from February 2018 to April 2021 (Fig. 1).
Fig. 1
Flowchart of the selection process for studies included in the systematic review.
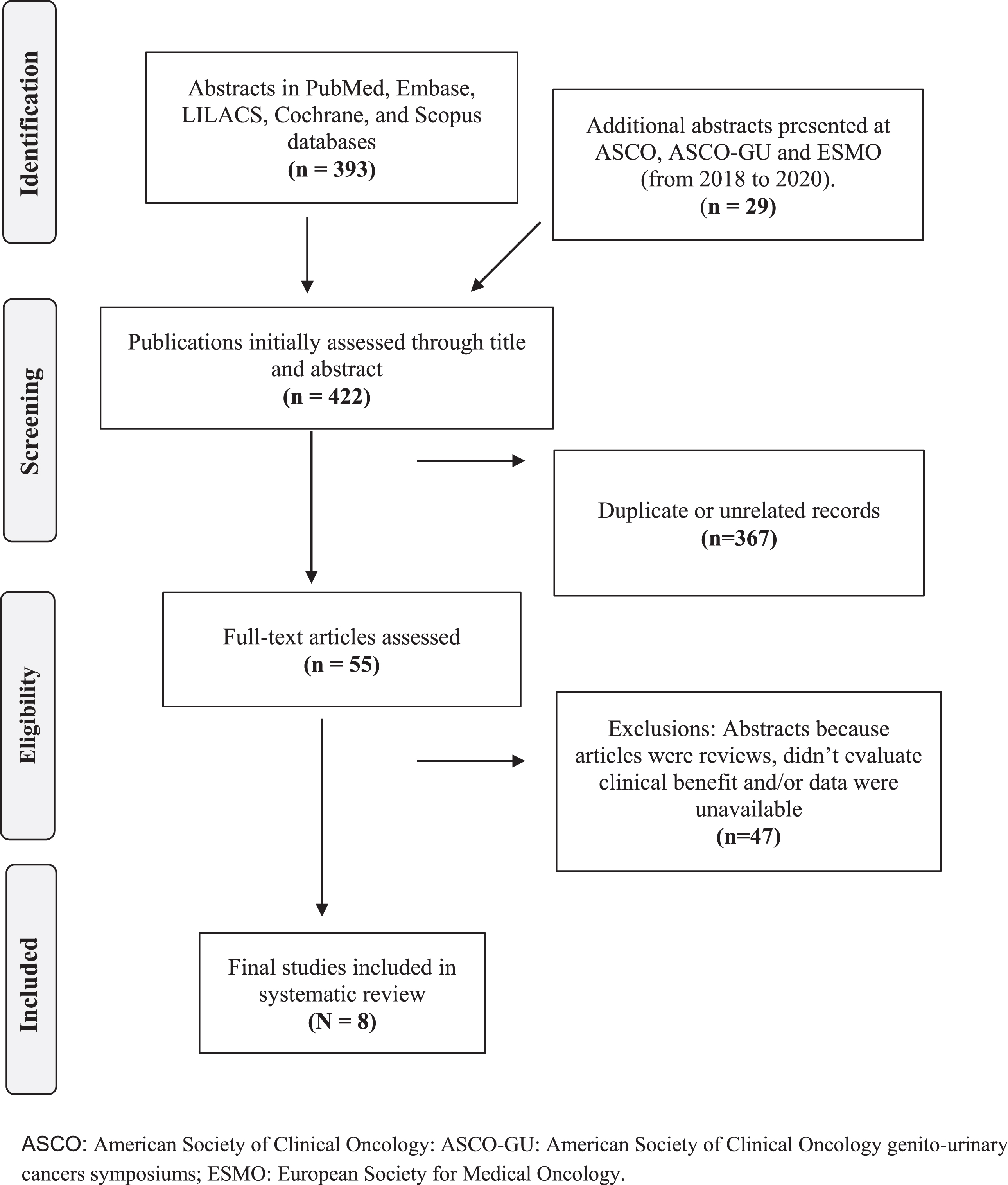
The inclusion criteria were as follows: (1) prospective and/or retrospective trials; (2) data on clear cell RCC; (3) English language. Trials without data of efficacy, published in non-peer-review journals, reviews and case reports were excluded.
Articles obtained from the research were selected independently by the three authors (APCDC, FSMM and AS), following the predetermined inclusion and exclusion criteria. Any differences in the selection of the article between the authors were resolved by discussion. Each stage of the research and the final result are summarized in Fig. 1.
Data extraction
The final abstracts selected had the following data extracted: number of patients, treatment protocol, presence or absence of PBRM1 mutation and efficacy data, which included overall response rate (ORR), progression-free survival (PFS), overall survival (OS) and CB, which was defined as complete response (CR), partial response (PR) or stable disease (SD) as defined by response evaluation criteria in solid tumors (RECIST) 1.1 (e.g., tumor shrinkage < 30% from baseline), for at least 6 months.
Outcome measures
The objective of this review was to evaluate a relationship between PBRM1 mutations, CB and efficacy with immunotherapy in patients with mRCC. Therefore, we sought to select for final analysis articles involving treatment with any of the ICI regimens, assessment of PBRM1 status (mutated or not mutated) and treatment efficacy.
Statistical analysis
The data extracted from the selected studies included in the final assessment were analyzed by the authors, taking into account the particularities and limitations of each one. No statistical analysis was conducted in this systematic review.
RESULTS
The systematic search identified 8 studies that met all inclusion criteria totaling 2029 patients with mRCC, the majority with clear cell renal cell carcinoma (ccRCC). The details of selected trials are summarized in Table 1.
Table 1
Summary of the Trials
| Author | Design | N | Biologic Sample | Method | Treatment Setting |
| Motzer R et al. [24] | Post-hoc analysis | 481 | FFPE tumor tissue | WES | First-line |
| McDermott et al. [23] | Post-hoc analysis | 262 | FFPE tumor tissue and peripheral blood | WES | First-line |
| Miao D et al. [25] | Prospective | 98 | FFPE tumor tissue and normal tissue | WES | ≥1 line |
| Braun D et al. [26] | Post-hoc analysis | 382 | FFPE tumor tissue | NI | ≥1 line |
| Braun D et al. [27] | Post-hoc analysis | 261 | FFPE tumor tissue and peripheral blood | WES | ≥1 line |
| Hakimi A et al. [16] | Retrospective | 189 | FFPE tumor tissue | TEs (MSK-IMPACT™) | ≥1 line |
| Vano YA et al. [29] | Retrospective | 324 | FFPE tumor tissue | IHC | ≥1 line |
| Dizman N et al. [30] | Retrospective | 32 | FFPE tumor tissue and Peripheral Blood | WES | ≥1 line |
FFPE: formalin-fixed, paraffin-embedded; IHC: immunohistochemistry; NI = not informed; MSK-IMPACT: Memorial Sloan Kettering - Integrated Mutation Profiling of Actionable Cancer Targets); TEs: transposable elements; WES: whole exome sequencing.
Genomic alterations in first-line treatment
Exploratory analysis of two prospective randomized controlled trials (RCT) evaluated the relationship between PBRM1 mutations and clinical outcomes in patients with metastatic predominantly ccRCC receiving ICI in the first-line treatment setting. The IMmotion150 trial randomized patients to receive atezolizumab + bevacizumab (atezo + beva) or atezolizumab (atezo) or sunitinib (sun) [22]. In a post-hoc analysis, aiming to identify molecular genomic alterations associated with overall response rate (ORR) and clinical outcomes, VHL and PBRM1 were the most frequent gene alterations in this population, found in 62% and 44% of the patients, respectively. PBRM1 mutations were associated with lower ORR in both ICI arms and were not associated with an improvement in the probability of progression-free survival (PFS). Interestingly, for patients who received sun, the authors found that PBRM1 mutations were associated with better PFS (HR 0.38; 95% CI, 0.20– 0.73) [23].
The second RCT was the CheckMate 214 study that randomized patients with metastatic ccRCC to ipilimumab + nivolumab (ipi + nivo) or sun [12]. In an exploratory analysis of this RCT, Motzer et al investigated possible biomarkers involved in the response to ipi + nivo. Using the samples of the pre-treatment tissue from enrolled patients, the authors conducted a search for the expression of PD-L1 by immunohistochemistry (IHC), genetic mutations by whole-exome sequencing (WES) and RNA sequencing. A total of 262 patients with mRCC who received ipi + nivo had their tumor genomic characteristics evaluated and 137 (52%) were PBRM1 mutated. There was neither OS nor PFS benefit derived from ICI in this specific subset of patients comparing with the PBRM1 wild-type patients. Unlike what was demonstrated in the IMmotion150 trial, in the CheckMate 214 study, the patients treated with suninitib did not show any benefit in PFS or OS in PBRM1 populations [24].
All the results in first-line treatment are summarized in Table 2.
Table 2
Studies assessing efficacy of first line ICI and PBRM1 mutational status in mRCC patients
| Author | N | Study | PBRM1 Mutation | Treatment | ORR (%) | CB (%) | PFS | OS |
| McDermott et al. [23] | 136 | IMmotion 150 | 1. Atezo: 30% x 41% | 3. Atezo | Atezo: ∼25% x ∼40% | 62% x 48% | 5. Atezo: 5.49 m x 8.15 m; HR: 1.33 (95% CI: 0.73– 2.42); p = 0.358 | |
| 2. Atezo + Beva: 29% x 36% | 4. Atezo + Beva | Atezo + Beva: ∼15% x ∼28% (p = 0.04) | 6. Atezo + Beva: 14.09 m x 8.31 m; HR: 0.67 (95% CI: 0.36– 1.25); p = 0.205 | NA | ||||
| Motzer RJ et al. [24] | 262 | CheckMate 214 | 137 x 125 | Nivo + Ipi | NA | NA | p = 0.76 | p = 0.11 |
Atezo: atezolizumab Beva: bevacizumab; CB: clinical benefit; CI: confidence interval; HR: hazard ratio; Ipi: ipilimumab; NA: data not available; Nivo: Nivolumab; ORR: overall response rate; OS: overall survival; PFS: progression-free survival.
Genomic alterations beyond first-line treatment
PBRM1 mutations were also evaluated in second and latter treatment lines in several studies. As part of a phase Ib prospective clinical trial, Miao et al. performed WES of mRCC from 35 patients and demonstrated superior clinical outcomes associated with LOF mutations in the PBRM1 gene (p = 0.012) [25]. This finding was confirmed in a parallel independent validation cohort of 63 ccRCC patients treated with ICI alone or in combination (p = 0.0071). Of note, the authors searched to hotspot mutations and PBRM1 was the only gene in which LOF mutations were enriched in the tumors from patients in the CB vs. non-CB (NCB) group. Patients whose tumors showed biallelic PBRM1 loss had significantly prolonged OS and PFS compared to patients without PBRM1 LOF (p = 0.0074 and p = 0.029, respectively) and they experienced sustained reductions in tumor burden. Patients who had no CB experienced progressive disease (PD), as determined by RECIST 1.1 and were discontinued from immunotherapy within 3 months. All other patients were termed “intermediate benefit” (IB). It is also important to highlight that tumors of patients with CB and no CB showed similar mutation burdens and intratumoral heterogeneity [25].
In the CheckMate 025 study, patients with mRCC who had progressed on VEGFR targeted therapy were randomized to receive nivolumab or everolimus [11]. In a post-hoc analysis of this RCT, Braun et al obtained tumor tissue from 382 of the 821 patients enrolled in the study and searched for putative truncated mutations (frameshift insertion/deletion, nonsense, splice location) in the PBRM1 gene. The mutation rate was 26.1% among all participants and the groups with or without mutations were homogeneous. Among patients with mutations in PBRM1, treatment with nivolumab resulted in modest but superior ORR (27% versus 17%) and CB (33% versus 25%), compared to wild-type patients. PFS (5.6 versus 2.9 months; HR, 0.67; 95% CI, 0.47– 0.96; p = 0.03) and OS (27.9 versus 20.9 months; HR, 0.65; 95% CI, 0.44– 0.96; p = 0.03) were also superior in patients with PBRM1 mutations. On the other hand, there was no benefit in patients with PBRM1 mutations treated with everolimus [26].
Hakimi et al, using 2 distinct cohorts with different types of cancer [the ICI database of the Memorial Sloan Kettering Cancer Center (MSKCC) and a cohort of 594 ccRCC patients with transcriptomic data], explored the effects of PBAF LOF in clinical results and TME. In the MSKCC cohort 40, 20, 38 and 3% of the 189 patients with ccRCC were treated with ICI as single-agent, ICI + ICI, ICI + VEGF and ICI + other treatments, respectively. Regarding the treatment setting, 51% received therapy as first-line and 49% as second or latter lines. In the entire mRCC cohort, presence of PBRM1 mutations in patients receiving ICI were neither associated to longer TTF nor superior OS, irrespectively of treatment line. Furthermore, in multivariate models adjusted for TMB and drug class, TTF was not significantly superior, as well as OS, when adjusted for IMDC risk score and line of therapy. Therefore, no association was found between favorable response to ICI and PBAF complex mutations [16].
Braun et al. selected patients from 3 prospective clinical trials with nivolumab in mRCC (CheckMate 009, CheckMate 010 and CheckMate 025) to perform an integrated genetic, transcriptomic and immunopathological analysis of their tumors. The WES data were available from 454 patients, of which 261 were treated with nivolumab and 193 with mTOR inhibition. VHL and PBRM1 mutations were the most frequent mutations, identified in 65 and 33% of the patients, respectively. Of note, most patients were previously treated with antiangiogenics. Pooled analysis of all three cohorts confirmed association of truncating mutations in PBRM1 with improved ORR, PFS and OS [27]. Data obtained from the CheckMate 009 trial indicated that truncating mutations in PBRM1 were associated with improved survival following PD-1 blockade in patients who progressed with antiangiogenic therapy, which was then validated using the CheckMate 025 cohort.
NIVOREN GETUG-AFU 26 is a prospective phase 2 trial with 720 patients that reported safety and efficacy of nivolumab in mRCC in a real-world scenario. All patients had failed to at least one line of VEGF/VEGFR inhibition and 21.4% had also received mTOR inhibitors [28]. As part of a translational research program, evaluating predictor biomarkers of response to ICI, the authors aimed to characterize immune cells into and around the tumor through IHC and associate these findings with clinical results obtained with nivolumab. The tumor samples of the patients were collected, reviewed and analyzed by IHC with a total of 13 markers, including PBRM-1, which was considered mutated when there was protein loss (absence of IHC staining). PBRM-1 loss was detected in 15% of the 324 patients included in the ancillary cohort and showed a trend in PFS and statistically improvement in OS [29].
On the other hand, in another retrospective trial with 91 patients, 38% with PBRM1 mutations and 35% treated with ICI (nivolumab + ipilimumab in first-line or nivo in second- or third-line setting) Dizman et al failed to show any association of PBRM1 mutation and CB with ICI (nivolumab or nivolumab + ipilimumab) [30].
All the results beyond first-line treatment are summarized in Table 3.
Table 3
Studies assessing efficacy of second-line ICI and PBRM1 mutational status in mRCC patients
| Author | Study | N | PBRM1 mutated x non-mutated | Treatment Protocol | ORR | CB | PFS | OS |
| Miao et al. [25] | Cohort 1: prospective (phase I Nivo trial) | 35 | 19 (54%) x 16 (46%) | Nivo | NE | 17/27 x 4/19 (OR: 6.10, 95% CI, 1.42– 32.64; p = 0.0071) | Cohort 1: p = 0.029 | Cohort 1: p = 0.0074 |
| Cohort 2: retrospective validation cohort | 63 | 28 (44%) x 35 (56%) | Anti PD-(L)1 + /- Anti CTLA-4 | Combined cohorts (98 pts): | ||||
| 1L (17 patients): p = 0.91 | ||||||||
| 2L and later lines (81 pts): p = 0.0087 | ||||||||
| Braun et al. [26] | CM-025 (post-hoc analysis) | 189 | 55 (29%) x 134 (71%) | Nivo | 27% x 17% (p = 0.04) | 62% x 48% | 5.6 m (3.6– 11.2) x 2.9m (2.0– 5.6) | 27.9m (19.9-NE) x 20.9m (18.3– 24.8) |
| HR:0.67 (95% CI, 0.47– 0.96); p = 0.03 | HR: 0.65 (95% CI: 0.44– 0.96); p = 0.03 | |||||||
| Hakimi et al. [16] | Retrospective data (MSKCC) | 189 | 32% x 14% | Anti PD-(L)1 (81.5%) and Anti CTLA-4 + Anti PD-(L)1 (18.5%) | NE | NE | Non statistical difference in the entire cohort (LOF HR: 0.73, p = 0.11; non-LOF HR: 1.05, p = 0.84), in first-line (HR: 0.6, p = 0.075) and second-line (HR: 0.87, p = 0.61) | NSD in the entire cohort (LOF HR: 1.5, p = 0.16; non-LOF HR: 1.05, p = 0.91), 1L (HR: 1.7, p = 0.29) and 2L (HR: 1.71, p = 0.44) |
| Braun DA et al. [27] | CM-009/010/025 (post-hoc analysis) | 261 | 85 (33%) x 176 (67%) | Nivo | — | Improve CB (p = 0.005) | p = 0.0056 | p = 0.00093 |
| Vano et al. [29] | NIVOREN GETUG-AFU26 | 323 | 49 (15%) x 274 (85%) | Nivo | — | — | 5.3 m x 3.9 m | 12m: 83.7% (70% – 91.5%) x 74% (68.4% – 78.8%) |
| HR: 0.75; p = 0.1 | HR: 0.5, p = 0.05 | |||||||
| Dizman et al. [30] | Retrospective data (COH) | 32 | 38% x 62% | Nivo + Ipi (1L) or Nivo (2L) | 22% (all ICI- treated patients) | p = 0.265 | 15.6 m (all ICI- treated patients) | — |
1L.: first-line; 2L: second-line; Atezo.: Atezolizumabe; CB: clinical benefit; CI: confidence interval; CM: CheckMate; COH: City of Hope; DFCI: Dana-Farber Cancer Institute; HR: hazard ratio; ICI: immune checkpoint inhibitors; Ipi: ipilimumab; LOF: loss of function; N: number of patients; NA: data not available; NE: not evaluated Nivo: nivolumab; NSD: no statistical difference; ORR: overall response rate; OS: overall survival; PFS: progression-free survival.
DISCUSSION
This review showed some interesting and controversial data about the association of genomic profiling of mRCC (specially PBRM1 mutations) and benefit with ICI treatment. Two of the trials, IMmotion150 and CheckMate 214, are pure first line reported trials and will be discussed separately. The exploratory molecular analyses from IMmotion150 not only failed to show association between PBRM1 mutations and CB with ICI, but data suggested instead superior clinical outcomes with sunitinib (the control arm) in this subset of patients. Considering only patients who received ICI, treatment with the combination of atezolizumab and bevacizumab, when confronted with atezolizumab monotherapy, showed improved PFS (HR 0.42; 95% CI, 0.22– 0.82). Angio gene signatures expression was higher in the VHL and PBRM1 mutant patients, which can explain better results with sunitinib [23]. This finding is in agreement with previously published data suggesting that PBRM1 mutations apparently enhance pro-angiogenic microenvironment of RCC, and consequently more sensible to TKI [31]. In an exploratory analysis of CheckMate 214 evaluating genomic biomarkers of response to ICI there was no benefit in OS or PFS according to PBRM1 mutation status [24]. These results are in concordance with pre-clinical data showing a cold TME with lower CD8 T infiltration, lower PD-L1 expression and higher expression of angiogenic gene signature in patients with PBRM1 mutations [17]. In this context, the cohort of COMPARZ trial showed that PBRM1 mutations in mRCC are associated with both higher angiogenic gene expression and response to anti-VEGF therapy, corroborating previous findings that links these mutations with superior TTF and PFS when treated with TT, as seen with sunitinib in the exploratory analysis of the IMmotion150 trial [23, 32].
IMmotion150 and CheckMate 214 analyses did not show superior efficacy of ICI in patients with PBRM1 mutations. The trials share similarities, such as the evaluation of treatment-naive metastatic patients with predominantly clear cell histology, the choice of sunitinib in the control arm, and the same efficacy outcomes (PFS, OS and ORR). Despite this, differences in the exploratory analysis of biomarkers and distinct levels of information provided regarding the molecular evaluation in each study contribute to the disparity between them. Although both studies adopted whole exome sequencing (WES) to detect mutations in PBRM1, in the IMmotion150 trial, the test was performed both on the tumor and peripheral blood mononuclear cells (PBMC), while in the CheckMate 214 study this information was not specified by authors. The IMmotion150 study, on the other hand, specifies that all of the 44% of the patients analyzed had LOF mutations of PBRM1, while in the CheckMate 214 study the authors did not mention whether the PBRM1 mutations detected in 52% of patients were LOF or non-LOF. The absence of this information can be explained by the fact that the CheckMate 214 study has not yet been published and the data available to date have been those presented in the oral presentation of ASCO 2020. Both studies lack data on the clonality of mutations in PBRM1, information that, based on previous studies, correlates with the response to ICI. Especially in melanoma and lung cancer an association was found between clonal mutations, clonal neoantigens and response to treatment, with tumors with clonal mutations being more likely to respond to ICI than those with subclonal mutations [15, 33].
The heterogeneity of renal cancer is a potential bias in these studies, since the molecular evaluation is usually performed from a single tumor sample. Thus, not all molecular alterations existing in the tumor will necessarily be present in the sample collected, and, as a result, existing genetic mutations may not be detected. The biopsy site of the tissue sample used for genetic evaluation also plays an important role and may impact on the molecular evaluation of the tumor. Tissue samples collected from metastatic sites may contain molecular and genomic changes that were acquired during the course of the disease and not present at the primary site of the tumor, impairing the evaluation result [27]. Finally, differences in the therapies can also be a reason, since IMmotion150 patients received atezolizumab and bevacizumab or just atezolizumab, while CheckMate 214 patients received a combination of ICI (nivolumab and ipilimumab).
The molecular analysis of patients enrolled in the JAVELIN Renal 101 trial and its association with the PFS result were evaluated by Motzer et al. [34]. The study was not found in our systematic search, so for this reason the data were not reported in our results. However, we chose to include it in the discussion due to the relevance of its findings, which are in line with those of the two other first-line studies just described. In their analysis, PBRM1 mutations were not associated with PFS improvement in both arms and OS data has not yet been published. Classical biomarkers such as PD-L1 and TMB were not able to predict response to ICI in patients with mRCC. In addition, data from an exploratory analysis suggest that antiangiogenic therapy plays an immunomodulatory role, a finding that may help to personalize therapeutic strategies for patients with different types of tumor. Similar to the other first-line trials, patients were predominantly of clear cell histology, with treatment-naive metastatic disease, the comparator arm was sunitinib, and the study outcomes were the same. Again, the clonality of the PBRM1 mutation has not been reported. The caveats regarding mutation analysis are the same as for CheckMate 214, since all data were analyzed using WES, but there are no details on the samples tested (primary tumor versus metastatic site) nor if the mutations detected in PBRM1 were LOF or non-LOF. It is important to note that the drug combination used in this trial is quite different, since the patients in the study arm received a combination of avelumab and axitinib.
Although mutations in PBRM1 do not appear to confer any benefit for metastatic patients with predominantly ccRCC receiving ICI as first-line therapy, there are technical limitations and differences between published articles, therefore additional studies should be carried out to clarify this issue.
While exploratory analysis of clinical trials in the first line setting failed to show benefit derived from ICI in patients harboring mutations in PBRM1, the data obtained in the second and subsequent lines of treatment have shown mixed results, with modestly superior clinical outcomes for these patients when exposed to ICI.
Of the 6 studies evaluating the benefit of ICI in tumors with PBRM1 mutations, 4 studies demonstrate benefit in these patients, and 2 studies are negative. At first, it appears that patients with PBRM1 mutations derived benefits from ICI, but all the studies have several caveats. The populations included in these studies are heterogeneous regarding to histological subtypes, lines of treatment, and the therapies used. The study of Hakimi et al [16] was the only one that included exclusively ccRCC patients. Dizman et al. [30] reported the impact of PBRM1 mutations in patients with both clear and non-clear cell histology. In this trial, 81% of patients had ccRCC, while 13%, 3%, and 3% had papillary, chromophobes and sarcomatoid tumors, respectively. The other studies evaluate therapies in patients with ccRCC predominantly, but without details about the other subtypes or the percentage of each one. The heterogeneity of histology is an important issue because the differences in the genomic landscape, in the genomic signature, in the TMB and in the infiltration of CD8 + T cells between clear and non-clear cell kidney tumors [15, 27].
Studies evaluating only the second- and later lines of treatment were those reported by Dizman et al. [30] and Vano et al. [29]. All other articles included patients in the first-line, second-line or beyond. The study by Miao et al has only 10% of patients in first-line therapy, while those of Hakimi et al. [16] has 51% of patients in first-line therapy. Efficacy data has not been reported according to the therapy line. It is important to highlight, as discussed above, the relationship between angiogenic signatures and PBRM1 mutations, and it is possible that patients who received antiangiogenic therapy may have some interference in the response to an ICI in the future. The only study that reported the results of the first- and second- or other treatment lines separately was that of Miao et al., in which the authors showed that patients with PBRM1 LOF mutations who received ICI in advanced lines reached superior PFS than patients who received it in the first line (p = 0.009).
In only 3 studies (Braun et al. [26, 27] and Vano et al. [29]) and in the primary cohort of Miao et al, patients received nivolumab as a single therapy. In the validation cohort of the study by Miao et al and in all others, the therapies were as follows: antiPD1/L1 + anti-CTLA-4, anti-PD1/L1 + anti-VEGF-TKI, anti-PD1/L1 + another drug or anti-CTLA-4 alone. The variety of therapeutic options and the different combinations used impact both the analysis of the efficacy of ICI as monotherapy and the evaluation of combined treatments, be it ICI or ICI associated with anti-VEGF-TKI.
Although all studies analyzed patients with PBRM1 LOF mutations, the different techniques used to detect these mutations is another issue. Three studies detected mutations through WES (Miao et al, Braun et al and Dizman et al), but only Miao et al matched the molecular analyses of the tumor samples with those of normal tissue. The other two studies compared the tumor sample with peripheral blood mononuclear cells (PBMC). Hakimi et al (MSK-IMPACT) used specific next-generation sequencing molecular panels from his respective institution to track mutations in PBRM1 and did not evaluate the materials in a paired manner. There is no report of which method Braun et al analyzed PBRM1 in one of their studies. Vano et al, in turn, investigated mutations in PBRM1 through immunohistochemistry (IHC).
Although data show greater sensitivity and specificity of IHC when compared to molecular tests by the Sanger method in the detection of LOF mutations in PBRM1, the comparison of this result with those of the other studies may be impaired [35]. As previously discussed, clonality appears to be important for increasing immunogenicity, but Miao et al were the only authors who described it, in their validation cohort. Issues about tissues are also relevant and when it comes to the large and well-established heterogeneity of kidney cancer, a single sample collected at a single site of the disease is unable to capture these differences. Another issue regarding tissue concerns the origin of the evaluated sample, whether from the primary tumor, from a synchronous metastasis, or from a metastatic site previously exposed to systemic therapy. In addition, the genomic landscape, immunogenicity, and gene signatures are not static factors and, when evaluated, do not necessarily represent the molecular characteristics of the disease at the time the patient received ICI.
In general, the study outcomes were homogeneous (OS, PFS, ORR and CB). The only trial that used a different definition of CB was that of Miao et al, who defined it as CR, PR and SD with some objective reduction in tumor burden, lasting at least 6 months. SD patients, but without objective reduction, were considered to be of intermediate benefit.
Another important bias is the heterogeneity of the studies, illustrated by the different methodologies (some were prospective and others retrospective) to the sizes of the populations, resulting in different levels of robustness and statistical confidence.
An important piece of information about PBRM1 non-LOF mutations would be the precise location where they occur. PBRM1 contains six bromodomains and is the only protein in the SWI/SNF complex that has more than one bromodomain. BD2 and BD4 seem to be the most important of them, and mutations in these sites can result in a greater impact on RCC biology and in response to therapies [18, 36, 37].
One trial excluded from results was by Alaiwi et al. [38] because the data regarding mRCC was available only in the presentation at ASCO Annual Meeting and not in the published abstract. In this trial, Alaiwi et al checked the association of ICI and PBRM1 mutation in patients treated at Dana-Farber Cancer Institute (DFCI). A cohort of 676 patients with 9 different histologic types of metastatic tumor (mainly non-small cell lung cancer, melanoma, RCC and bladder cancer) showing any truncation mutation or homozygous deletion in one of the PBAF-complex genes and who received ICI in the metastatic setting was selected. The sample size of patients with mRCC was 68 and an association with mutations in PBAF-complex and improved OS (not reached versus 10.9 months; p = 0.004) and time to treatment failure (TTF) (11.3 versus 5.6 months; p = 0.017) was observed. The ORR was numerically higher in the group with mutation (32 versus 17%; p = 0.259) but without statistical significance. CB was statistically improved in the patients with mutations (71% versus 40%; p = 0.014). This trial shares the same weakness from the others, as well as no data about the histology of the tumors, almost half of patients in first-line treatment, different ICI used, alone or in combination, PBRM1 mutation status was tested using a next-generation sequencing (NSG) from the author’s institution (Oncopanel/DFCI) with no paired sample, no clonality describe, the retrospective nature of the study, a small population, and all issues regarding the tumor samples [38].
Although conflicting data have been presented connecting PBRM1 LOF mutations with superior clinical outcomes with ICI in mRCC patients, this review suggests that such genomic alterations play a role, particularly in the advanced lines of treatment (second line onwards). One possible explanation for this difference across lines of treatment is that most patients in advanced lines have been previously exposed to antiangiogenics, which could influence the TME turning it more immunologic active (“hot”) and, therefore, more favorable to ICI activity [39, 40].
It is important to emphasize the heterogeneity of the studies in this systematic review and its impact. Differences in histology, treatment lines, choice of drugs, molecular assessment techniques, tissue issues, definitions of PBRM1 mutations, distinct clinical results and the different nature of the trials represent significant confounding factors in the interpretation of the data presented here.
Considering future directions regarding PBRM1 mutations as a potential predictive biomarker for immunotherapy in mRCC, there is a huge potential for clinical trials. Currently, there are no prospective clinical trials with that purpose, and only one prospective observational clinical trial evaluating a prediction model in localized RCC using molecular biomarkers, including PBRM1 mutations [41]. Regarding the role of other potential biomarkers of clinical response to ICI in RCC, PD-L1 expression remains controversial as the results obtained in the exploratory analyses of the landmark clinical trials on that matter are inconclusive [11– 14, 22]. The data on TMB and its potential to predict response to immunotherapy in the context of mRCC are conflicting. Although the exploratory analysis of IMmotion150 has not shown an association between TMB and superior clinical benefit with nivolumab, post-hoc analysis of NIVES, an ongoing phase 2 study that investigates the combination of nivolumab with stereotactic body radiotherapy in patients pretreated with mRCC, can help to clarify this issue [42]. RCC typically features a variety of tumor-infiltrating immune system cells. The different subpopulations of immune cells and the diverse possibilities of interaction between them can influence the tumor’s sensitivity to ICIs. Specific genomic signatures, based on gene expression profiles predictive of immune system activation and antitumor response, were investigated as predictive biomarkers of response to immunotherapy. Exploratory analysis of IMmotion150 reported an association between a T-effector immune gene signature, PD-L1 expression and tumor infiltration of CD8 + T cells, implying greater ORR and PFS with atezolizumab [23]. In general, however, conflicting data have been published so far and further studies on this topic are awaited.
A possible correlation between the development of immune-related adverse events (irAEs) and superior outcomes with ICIs has also been studied across different types of tumors. In patients with mRCC, data from treatment with nivolumab in the second line or beyond showed that those who manifested irAEs obtained significantly longer OS [43]. Retrospective data from Asian patients with mRCC receiving nivolumab corroborate these findings [44]. Despite being provocative data, a number of unanswered questions still remain, such as whether any type of irAEs can correlate with a better response to ICI or whether only specific types trigger this mechanism.
One of the weaknesses of conventional transcriptomic methods is that they do not account for the heterogeneity of tumor cells, since they are usually carried out in bulk and, therefore, their data represent an average of gene expression of several cells. Thus, NGS-based technologies are now increasingly focused on the evaluation of cells at the individual level and the sequencing of single cell RNA (scRNA-seq) has overcome this problem. The analysis of the gene expression of individual cells, exploring their differences, may allow the identification of distinct and even unknown cell populations, a leap in the potential to impact our understanding of phenomena such as resistance to cancer treatments and relapses. This technology could even be incorporated into the analysis of circulating tumor cells (CTCs) in the blood, also known as “liquid biopsy”. The combination of the two methods would not only detect CTCs, but also track coding mutations and fusion genes [42, 45].
CONCLUSIONS
This systematic review showed an association between truncated PBRM1 mutations and superior efficacy with ICI in patients with mRCC predominantly clear cell histology, receiving nivolumab as second- or later treatment lines, but not in those in the first-line setting. Although many patients with mRCC with LOF mutations in PBRM1 have the greatest clinical benefit with ICI, the data published so far are conflicting. Clinical validation in a large prospective study is necessary before PBRM1 mutations can be considered a predictive genomic tool for use in daily clinical practice.
ACKNOWLEDGMENTS
The authors have no acknowledgements.
FUNDING
The authors report no funding.
AUTHORS CONTRIBUTIONS
All authors contributed equally in all stages of the article’s development (conception, execution and interpretation of data).
CONFLICT OF INTEREST
André Paternò Castello Dias Carneiro: no conflicts of interest. Fernando Sabino Marques Monteiro Speaker: MSD, Janssen, Astellas and BMS. Advisory Board: Janssen, MSD and BMS. Research Funding: Janssen. Founder/Owner: BIO (Brazilian Information Oncology). Andrey Soares: Financial - Honoraria: Janssen, Pfizer, Bayer, Novartis, AstraZeneca, Astellas Pharma, Pierre Fabre, Merck Serono, Sanofi, Roche and Ipsen. Consulting or Advisory Role: Astellas Pharma, Janssen, Roche, Bayer, Lilly, AstraZeneca, Novartis, MSD, Bristol-Myers Squibb, Zodiac, Amgem, Ipsen and United. Research Funding: Bristol- Myers Squibb (Inst). Travel, Accommodations, Expenses: AstraZeneca, Pfizer, AstellasPharma, Bristol-Myers Squibb, Bayer, Roche, Janssen, Merck Serono, Sanofi and Ipsen. Nonfinancial – No other potential conflicts of interest were reported. Founder/Owner: BIO (Brazilian Information Oncology).
REFERENCES
[1] | Robert C . A decade of immune-checkpoint inhibitors in cancer therapy. Nat Commun. (2020) ;11: :3801. https://doi.org/10.1038/s41467-020-17670-y. |
[2] | Davis AA , Patel VG . The role of PD-L1 expression as a predictive biomarker: an analysis of all US Food and Drug Administration (FDA) approvals of immune checkpoint inhibitors. J Immunother Cancer. (2019) ;7: (1):278. doi: 10.1186/s40425-019-0768-9. PMID: 31655605; PMCID: PMC6815032. |
[3] | Marabelle A , Le DT , Ascierto PA , Di Giacomo AM , De Jesus-Acosta A , Delord J-P , Geva R , Gottfried M , Penel N , Hansen RA , Piha-Paul AS , Doi T , Gao B , Chung CH , Martin-Lopez J , Yung-Jue Bang , Frommer SR , Shah M , Ghori R , Joe KA , Pruitt KS , Diaz Jr, LA , Efficacy of Pembrolizumab in Patients With Noncolorectal High Microsatellite Instability/Mismatch Repair–Deficient Cancer: Results From the Phase II KEYNOTE-158 Study. Journal of Clinical Oncology. (2020) ;38: (1):1–10. |
[4] | Marabelle A , Fakih M , Lopez J , Shah M , Shapira-Frommer R , Nakagawa K , Chung HC , Kindler HL , Lopez-Martin JA , Miller WH Jr , Italiano A , Kao S , Piha-Paul SA , Delord JP , McWilliams RR , Fabrizio DA , Aurora-Garg D , Xu L , Jin F , Norwood K , Bang YJ . Association of tumour mutational burden with outcomes in patients with advanced solid tumours treated with pembrolizumab: prospective biomarker analysis of the multicohort, open-label, phase 2 KEYNOTE-158 study. Lancet Oncol. (2020) ;21: (10):1353–65. doi: 10.1016/S1470-2045(20)30445-9. Epub 2020 Sep 10. PMID: 32919526. |
[5] | Snyder A , Makarov V , Merghoub T , Yuan J , Zaretsky JM , Desrichard A , Walsh LA , Postow MA , Wong P , Ho TS , Hollmann TJ , Bruggeman C , Kannan K , Li Y , Elipenahli C , Liu C , Harbison CT , Wang L , Ribas A , Wolchok JD , Chan TA . Genetic basis for clinical response to CTLA-4 blockade in melanoma. N Engl J Med. (2014) ;371: (23):2189–99. doi: 10.1056/NEJMoa1406498. Epub 2014 Nov 19. Erratum in: N Engl J Med. 2018;379(22):2185. PMID: 25409260; PMCID: PMC4315319. |
[6] | Rizvi NA , Hellmann MD , Snyder A , Kvistborg P , Makarov V , Havel JJ , Lee W , Yuan J , Wong P , Ho TS , Miller ML , Rekhtman N , Moreira AL , Ibrahim F , Bruggeman C , Gasmi B , Zappasodi R , Maeda Y , Sander C , Garon EB , Merghoub T , Wolchok JD , Schumacher TN , Chan TA . Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science. (2015) ;348: (6230):124–8. doi: 10.1126/science.aaa1348. Epub 2015 Mar 12. PMID: 25765070; PMCID: PMC4993154. |
[7] | Le DT , Uram JN , Wang H , Bartlett BR , Kemberling H , Eyring AD , Skora AD , Luber BS , Azad NS , Laheru D , Biedrzycki B , Donehower RC , Zaheer A , Fisher GA , Crocenzi TS , Lee JJ , Duffy SM , Goldberg RM , de la Chapelle A , Koshiji M , Bhaijee F , Huebner T , Hruban RH , Wood LD , Cuka N , Pardoll DM , Papadopoulos N , Kinzler KW , Zhou S , Cornish TC , Taube JM , Anders RA , Eshleman JR , Vogelstein B , Diaz LA Jr. . PD-1 Blockade in Tumors with Mismatch-Repair Deficiency. N Engl J Med. (2015) ;372: (26):2509–20. doi: 10.1056/NEJMoa1500596. Epub 2015 May 30. PMID: 26028255; PMCID: PMC4481136. |
[8] | Van Allen EM , Miao D , Schilling B , Shukla SA , Blank C , Zimmer L , Sucker A , Hillen U , Foppen MHG , Goldinger SM , Utikal J , Hassel JC , Weide B , Kaehler KC , Loquai C , Mohr P , Gutzmer R , Dummer R , Gabriel S , Wu CJ , Schadendorf D , Garraway LA . Genomic correlates of response to CTLA-4 blockade in metastatic melanoma. Science. (2015) ;350: (6257):207–211. doi: 10.1126/science.aad0095. Epub 2015 Sep 10. Erratum in: Science. 2015;350(6262):aad 8366. Erratum in: Science. 2016 Apr 15;352(6283). pii: aaf8264. doi: 10.1126/science.aaf8264. PMID: 26359337; PMCID: PMC5054517. |
[9] | Cancer Genome Atlas Research Network. Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature. (2013) ;499: (7456):43–9. doi: 10.1038/nature12222. Epub 2013 Jun 23. PMID: 23792563; PMCID: PMC3771322. |
[10] | de Velasco G , Miao D , Voss MH , Hakimi AA , Hsieh JJ , Tannir NM , Tamboli P , Appleman LJ , Rathmell WK , Van Allen EM , Choueiri TK . Tumor Mutational Load and Immune Parameters across Metastatic Renal Cell Carcinoma Risk Groups. Cancer Immunol Res. (2016) ;4: (10):820–2. doi: 10.1158/2326-6066.CIR-16-0110. Epub 2016 Aug 18. PMID: 27538576; PMCID: PMC5050137. |
[11] | Motzer RJ , Escudier B , McDermott DF , George S , Hammers HJ , Srinivas S , Tykodi SS , Sosman JA , Procopio G , Plimack ER , Castellano D , Choueiri TK , Gurney H , Donskov F , Bono P , Wagstaff J , Gauler TC , Ueda T , Tomita Y , Schutz FA , Kollmannsberger C , Larkin J , Ravaud A , Simon JS , Xu LA , Waxman IM , Sharma P ; CheckMate 025 Investigators. Nivolumab versus Everolimus in Advanced Renal-Cell Carcinoma. N Engl J Med. (2015) ;373: (19):1803–13. doi: 10.1056/NEJMoa1510665. Epub 2015 Sep 25. PMID: 26406148; PMCID: PMC5719487. |
[12] | Motzer RJ , Tannir NM , McDermott DF , Arén Frontera O , Melichar B , Choueiri TK , Plimack ER , Barthélémy P , Porta C , George S , Powles T , Donskov F , Neiman V , Kollmannsberger CK , Salman P , Gurney H , Hawkins R , Ravaud A , Grimm MO , Bracarda S , Barrios CH , Tomita Y , Castellano D , Rini BI , Chen AC , Mekan S , McHenry MB , Wind-Rotolo M , Doan J , Sharma P , Hammers HJ , Escudier B ; CheckMate 214 Investigators. Nivolumab plus Ipilimumab versus Sunitinib in Advanced Renal-Cell Carcinoma. N Engl J Med. (2018) ;378: (14):1277–90. doi: 10.1056/NEJ-Moa1712126. Epub 2018 Mar 21. PMID: 29562145; PMCID: PMC5972549. |
[13] | Rini BI , Plimack ER , Stus V , Gafanov R , Hawkins R , Nosov D , Pouliot F , Alekseev B , Soulières D , Melichar B , Vynnychenko I , Kryzhanivska A , Bondarenko I , Azevedo SJ , Borchiellini D , Szczylik C , Markus M , McDermott RS , Bedke J , Tartas S , Chang YH , Tamada S , Shou Q , Perini RF , Chen M , Atkins MB , Powles T ; KEYNOTE-426 Investigators. Pembrolizumab plus Axitinib versus Sunitinib for Advanced Renal-Cell Carcinoma. N Engl J Med. . (2019) ;380: (12):1116–27. doi: 10.1056/NEJMoa1816714. Epub 2019 Feb 16. PMID: 30779529. |
[14] | Motzer RJ , Penkov K , Haanen J , Rini B , Albiges L , Campbell MT , Venugopal B , Kollmannsberger C , Negrier S , Uemura M , Lee JL , Vasiliev A , Miller WH Jr , Gurney H , Schmidinger M , Larkin J , Atkins MB , Bedke J , Alekseev B , Wang J , Mariani M , Robbins PB , Chudnovsky A , Fowst C , Hariharan S , Huang B , di Pietro A , Choueiri TK . Avelumab plus Axitinib versus Sunitinib for Advanced Renal-Cell Carcinoma. N Engl J Med. (2019) ;380: (12):1103–15. doi: 10.1056/NEJMoa1816047. Epub 2019 Feb 16. PMID: 30779531; PMCID: PMC6716603. |
[15] | Keenan TE , Burke KP , Van Allen EM . Genomic correlates of response to immune checkpoint blockade. Nat Med. (2019) ;25: (3):389–402. doi: 10.1038/s41591-019-0382-x. Epub 2019 Mar 6. PMID: 30842677; PMCID: PMC6599710. |
[16] | Hakimi A , Ari, Kyrollis Attalla , DiNatale Renzo G. , Ostrovnaya Irina , Flynn Jessica , Blum Kyle A. , Ged Yasser et al. “A Pan-Cancer Analysis of PBAF Complex Mutations and Their Association with Immunotherapy Response.”. Nature Communications. (2020) ;11: (1). Nature Research. doi:10.1038/s41467-020-17965-0. |
[17] | Liu XD , Kong W , Peterson CB , McGrail DJ , Hoang A , Zhang X , Lam T , Pilie PG , Zhu H , Beckermann KE , Haake SM , Isgandrova S , Martinez-Moczygemba M , Sahni N , Tannir NM , Lin SY , Rathmell WK , Jonasch E . PBRM1 loss defines a nonimmunogenic tumor phenotype associated with checkpoint inhibitor resistance in renal carcinoma. Nat Commun. (2020) ;11: (1):2135. doi: 10.1038/s41467-020-15959-6. PMID: 32358509; PMCID: PMC7195420. |
[18] | Cai W , Su L , Liao L , Liu ZZ , Langbein L , Dulaimi E , Testa JR , Uzzo RG , Zhong Z , Jiang W , Yan Q , Zhang Q , Yang H . PBRM1 acts as a p53 lysine-acetylation reader to suppress renal tumor growth. Nat Commun. (2019) ;10: (1):5800. doi: 10.1038/s41467-019-13608-1. PMID: 31863007; PMCID: PMC6925188. |
[19] | Sobhani N , D’Angelo A , Wang X , Young KH , Generali D , Li Y . Mutant p53 as an Antigen in Cancer Immunotherapy. Int J Mol Sci. (2020) ;21: (11):4087. doi: 10.3390/ijms21114087. PMID: 32521648; PMCID: PMC7312027. |
[20] | Blagih J , Buck MD , Vousden KH . p53, cancer and the immune response. J Cell Sci. (2020) ;133: (5):jcs237453. doi: 10.1242/jcs.237453. PMID: 32144194. |
[21] | Moher D , Shamseer L , Clarke M , Ghersi D , Liberati A , Petticrew M , Shekelle P , Stewart LA ; PRISMA-P Group. Preferred reporting items for systematic review and meta-analysis protocols (PRISMA-P) 2015 statement. Syst Rev. (2015) ;4: (1):1. doi: 10.1186/2046-4053-4-1. PMID: 25554246; PMCID: PMC4320440. |
[22] | Rini BI , Powles T , Atkins MB , Escudier B , McDermott DF , Suarez C , Bracarda S , Stadler WM , Donskov F , Lee JL , Hawkins R , Ravaud A , Alekseev B , Staehler M , Uemura M , De Giorgi U , Mellado B , Porta C , Melichar B , Gurney H , Bedke J , Choueiri TK , Parnis F , Khaznadar T , Thobhani A , Li S , Piault-Louis E , Frantz G , Huseni M , Schiff C , Green MC , Motzer RJ . Atezolizumab plus bevacizumab versus sunitinib in patients with previously untreated metastatic renal cell carcinoma (IMmotion151): a multicentre, open-label, phase 3, randomised controlled trial. The Lancet. (2019) ;393: (10189):2404–15. https://doi.org/10.1016/S0140-6736(19)30723-8. |
[23] | McDermott DF , Huseni MA , Atkins MB , Motzer RJ , Rini BI , Escudier B , Fong L , Joseph RW , Pal SK , Reeves JA , Sznol M , Hainsworth J , Rathmell WK , Stadler WM , Hutson T , Gore ME , Ravaud A , Bracarda S , Suárez C , Danielli R , Gruenwald V , Choueiri TK , Nickles D , Jhunjhunwala S , Piault-Louis E , Thobhani A , Qiu J , Chen DS , Hegde PS , Schiff C , Fine GD , Powles T . Clinical activity and molecular correlates of response to atezolizumab alone or in combination with bevacizumab versus sunitinib in renal cell carcinoma. Nat Med. (2018) ;24: (6):749–57. doi: 10.1038/s41591-018-0053-3. Epub 2018 Jun 4. Erratum in: Nat Med. 2018;24(12):1941. PMID: 29867230; PMCID: PMC6721896. |
[24] | Motzer RJ , Choueiri TK , McDermott DF , Yao J , Ammar R , Pappillon-Cavanagh S , Saggi SS , McHenry BM , Ross-Macdonald P , Wind-Rotolo M , et al. Biomarker analyses from the phase III CheckMate 214 trial of nivolumab plus ipilimumab (N+I) or sunitinib (S) in advanced renal cell carcinoma (aRCC). J Clin Oncol. (2020) ;38: :5009. |
[25] | Miao D , Margolis CA , Gao W , Voss MH , Li W , Martini DJ , Norton C , Bossé D , Wankowicz SM , Cullen D , Horak C , Wind-Rotolo M , Tracy A , Giannakis M , Hodi FS , Drake CG , Ball MW , Allaf ME , Snyder A , Hellmann MD , Ho T , Motzer RJ , Signoretti S , Kaelin WG Jr , Choueiri TK , Van Allen EM . Genomic correlates of response to immune checkpoint therapies in clear cell renal cell carcinoma. Science. (2018) ;359: (6377):801–6. doi: 10.1126/science.aan5951. Epub 2018 Jan 4. PMID: 29301960; PMCID: PMC6 035749. |
[26] | Braun DA , Ishii Y , Walsh AM , Van Allen EM , Wu CJ , Shukla SA , Choueiri TK . Clinical Validation of PBRM1 Alterations as a Marker of Immune Checkpoint Inhibitor Response in Renal Cell Carcinoma. JAMA Oncol. . (2019) ;5: (11):1631–3. doi: 10.1001/jamaoncol.2019.3158. PMID: 31486842; PMCID: PMC6735411. |
[27] | Braun DA , Hou Y , Bakouny Z , Ficial M , Sant’ Angelo M , Forman J , Ross-Macdonald P , Berger AC , Jegede OA , Elagina L , Steinharter J , Sun M , Wind-Rotolo M , Pignon JC , Cherniack AD , Lichtenstein L , Neuberg D , Catalano P , Freeman GJ , Sharpe AH , McDermott DF , Van Allen EM , Signoretti S , Wu CJ , Shukla SA , Choueiri TK . Interplay of somatic alterations and immune infiltration modulates response to PD-1 blockade in advanced clear cell renal cell carcinoma. Nat Med. (2020) ;26: (6):909–18. doi: 10.1038/s41591-020-0839-y. Epub 2020 May 29. PMID: 32472114; PMCID: PMC7499153. |
[28] | Albiges L , Negrier S , Dalban C , Chevreau C , Gravis G , Oudard S , Laguerre B , Barthelemy P , Borchiellini D , Gross-Goupil M , Geoffrois L , Rolland F , Thiery-Vuillemin A , Joly FLadoire S , Tantot F , Escudier B , “Safety and Efficacy of Nivolumab in Metastatic Renal Cell Carcinoma (MRCC): Final Analysis from the NIVORENGETUGAFU 26 Study.” Journal of Clinical Oncology 37 (7 suppl).American Society of Clinical Oncology (ASCO). 2019;542–542. doi:10.1200/jco.2019.37.7_suppl.542. |
[29] | Vano Y-A , Rioux-Leclercq N , Dalban C , Sautes-Fridman C , Bougoüin A , Chaput N ,... Albiges L . NIVOREN GETUG-AFU 26 translational study: Association of PD-1, AXL, and PBRM-1 with outcomes in patients (pts) with metastatic clear cell renal cell carcinoma (mccRCC) treated with nivolumab (N). Journal of Clinical Oncology. (2020) ;38: (6_suppl):618–618. https://doi.org/10.1200/jco.2020.38.6_suppl.618 |
[30] | Dizman N , Lyou Y , Salgia N , Bergerot PG , Hsu J , Enriquez D , Izatt T , Trent JM , Byron S , Pal S . Correlates of clinical benefit from immunotherapy and targeted therapy in metastatic renal cell carcinoma: comprehensive genomic and transcriptomic analysis. Journal for Immunotherapy of Cancer. (2020) ;8: (2):e000953. https://doi.org/10.1136/jitc-2020-000953 |
[31] | Voss MH , Kuo F , Chen D , Marker M , Patel P , Redzematovic A ,... Hakimi AA . Integrated biomarker analysis for 412 renal cell cancer (RCC) patients (pts) treated on the phase 3 COMPARZ trial: Correlating common mutation events in PBRM1 and BAP1 with angiogenesis expression signatures and outcomes on tyrosine kinase inhibitor (TKI) therapy. Journal of Clinical Oncology. (2017) ;35: (15_suppl):4523–4523. https://doi.org/10.1200/jco.2017.35.15_suppl.4523 |
[32] | Carlo MI , Manley B , Patil S , Woo KM , Coskey DT , Redzematovic A , Arcila M , Ladanyi M , Lee W , Chen YB , Lee CH , Feldman DR , Hakimi AA , Motzer RJ , Hsieh JJ , Voss MH . Genomic Alterations and Outcomes with VEGF-Targeted Therapy in Patients with Clear Cell Renal Cell Carcinoma. Kidney Cancer. (2017) ;1: (1):49–56. doi: 10.3233/KCA-160003. PMID: 30334004; PMCID: PMC6179122. |
[33] | Lu T , Wang S , Xu L , Zhou Q , Singla N , Gao J , Manna S , Pop L , Xie Z , Chen M , Luke JJ , Brugarolas J , Hannan R , Wang T . Tumor neoantigenicity assessment with CSiN score incorporates clonality and immunogenicity to predict immunotherapy outcomes. Sci Immunol.eaaz. (2020) ;5: (44):3199. doi: 10.1126/sciimmunol.aaz3199. PMID: 32086382; PMCID: PMC7239327. |
[34] | Motzer RJ , Robbins PB , Powles T , Albiges L , Haanen JB , Larkin J , Mu XJ , Ching KA , Uemura M , Pal SK , Alekseev B , Gravis G , Campbell MT , Penkov K , Lee JL , Hariharan S , Wang X , Zhang W , Wang J , Chudnovsky A , di Pietro A , Donahue AC , Choueiri TK . Avelumab plus axitinib versus sunitinib in advanced renal cell carcinoma: biomarker analysis of the phase 3 JAVELIN Renal 101 trial. Nat Med. (2020) ;26: (11):1733–1741. doi: 10.1038/s41591-020-1044-8. Epub 2020 Sep 7. PMID: 32895571. |
[35] | Ho TH , Kapur P , Joseph RW , Serie DJ , Eckel-Passow JE , Parasramka M , Cheville JC , Wu KJ , Frenkel E , Rakheja D , Stefanius K , Brugarolas J , Parker AS . Loss of PBRM1 and BAP1 expression is less common in non-clear cell renal cell carcinoma than in clear cell renal cell carcinoma. Urol Oncol. (2015) ;33: (1):23.e9–23.e14. doi: 10.1016/j.urolonc.2014.10.014. Epub 2014 Nov 24. PMID: 25465300; PM CID: PMC4274200. |
[36] | Conway J , Taylor-Weiner A , Braun D , Bakouny Z , Choueiri TK , van Allen EM . (2020, November 4). PBRM1 loss-of-function mutations and response to immune checkpoint blockade in clear cell renal cell carcinoma. MedRxiv. medRxiv. https://doi.org/10.1101/2020.10.30.20222356 |
[37] | Porter EG , Dykhuizen EC . Individual Bromodomains of Polybromo-1 Contribute to Chromatin Association and Tumor Suppression in Clear Cell Renal Carcinoma. J Biol Chem. (2017) ;292: (7):2601–10. doi: 10.1074/jbc.M116.746875. Epub 2017 Jan 4. PMID: 28053089; PMCID: PMC 5314159. |
[38] | Abou Alaiwi S , Nassar A , El Bakouny Z , Berchuck JE , Nuzzo P , Flippot R ,... Choueiri TK . Association of polybromo-associated BAF (PBAF) complex mutations with overall survival (OS) in cancer patients (pts) treated with checkpoint inhibitors (ICIs). Journal of Clinical Oncology. (2019) ;37: (15_suppl):103–103. https://doi.org/10.1200/jco.2019.37.15_suppl.103 |
[39] | Brodaczewska KK , Szczylik C , Kieda C . Immune consequences of anti-angiogenic therapyin renal cell carcinoma. Contemporary Oncology (Poznan, Poland).. (2018) ;22: (1A):14–22. DOI: 10.5114/wo.2018.73878. |
[40] | Tan HY , Wang N , Lam W , Guo W , Feng Y , Cheng YC . Targeting tumour microenvironment by tyrosine kinase inhibitor. Molecular cancer. (2018) ;17: (1):43. https://doi.org/10.1186/s12943-018-0800-6 |
[41] | Park JS , Pierorazio PM , Lee JH , Lee HJ , Lim YS , Jang WS , Kim J , Lee SH , Rha KH , Cho NH , Ham WS . Gene Expression Analysis of Aggressive Clinical T1 Stage Clear Cell Renal Cell Carcinoma for Identifying Potential Diagnostic and Prognostic Biomarkers. Cancers (Basel).. (2020) ;12: (1):222. doi: 10.3390/cancers12010222. PMID: 31963294; PMCID: PMC7017065. |
[42] | Masini C , Iotti C , De Giorgi U , Bellia RS , Buti S , Salaroli F ,... Pinto C . Nivolumab (NIVO) in combination with stereotactic body radiotherapy (SBRT) in pretreated patients (pts) with metastatic renal cell carcinoma (mRCC): First results of phase II NIVES study. Journal of Clinical Oncology. (2020) ;38: (6_suppl):613–613. https://doi.org/10.1200/jco.2020.38.6 suppl.613 |
[43] | Ishihara H , Takagi T , Kondo T , Homma C , Tachibana H , Fukuda H , Yoshida K , Iizuka J , Kobayashi H , Okumi M , Ishida H , Tanabe K . Association between immune-related adverse events and prognosis in patients with metastatic renal cell carcinoma treated with nivolumab. Urol Oncol. (2019) ;37: (6):355.e21–355.e29. doi: 10.1016/j.urolonc.2019.03.003. Epub 2019 Mar 29. PMID: 30935847. |
[44] | Kato R , Ikarashi D , Matsuura T , Maekawa S , Kato Y , Kanehira M , Takata R , Tokuyama R , Tamai K , Harigai N , Nakazaki Y , Obara W . Analyses of Nivolumab Exposure and Clinical Safety Between 3-mg/kg Dosing and 240-mg Flat Dosing in Asian Patients with Advanced Renal Cell Carcinoma in the Real-World Clinical Setting. Transl Oncol. (2020) ;13: (6):100771. doi: 10.1016/j.tranon.2020.100771. Epub 2020 May 3. PMID: 32375081; PMCID: PMC7205763. |
[45] | Olsen TK , Baryawno N . Introduction to Single-Cell RNA Sequencing. Curr Protoc Mol Biol. (2018) ;122: (1):e57. doi: 10.1002/cpmb.57. PMID: 29851283. |




