BRCA1-Associated Protein 1 (BAP-1) as a Prognostic and Predictive Biomarker in Clear Cell Renal Cell Carcinoma: A Systematic Review
Abstract
BACKGROUND:
The gene that encodes BRCA1-associated protein 1 (BAP1) has been reported to be dysregulated in several human cancers such as uveal melanoma, malignant pleural mesothelioma, hepatocellular carcinoma, thymic epithelial tumors, and clear-cell renal cell carcinoma (ccRCC). The gene is located on the human chromosome 3p21.3, encoding a deubiquitinase and acts as a classic two-hit tumor suppressor gene. BAP1 predominantly resides in the nucleus, where it interacts with several chromatin-associated factors, as well as regulates calcium signaling in the cytoplasm. As newer therapies continue to evolve for the management of RCC, it is important to understand the role of BAP1 mutation as a prognostic and predictive biomarker.
OBJECTIVE:
We aimed to systematically evaluate the role of BAP1 mutations in patients with RCC in terms of its impact on prognosis and its role as a predictive biomarker.
METHODS:
Following PRISMA guidelines, we performed a systematic literature search using PubMed and Embase through March 2021. Titles and abstracts were screened to identify articles for full-text and then a descriptive review was performed.
RESULTS:
A total of 490 articles were initially identified. Ultimately 71 articles that met our inclusion criteria published between 2012–2021 were included in the analysis. Data were extracted and organized to reflect the role of BAP1 alterations as a marker of prognosis as well as a marker of response to treatments, such as mTOR inhibitors, VEGF tyrosine kinase inhibitors, and immune checkpoint inhibitors.
CONCLUSIONS:
Alterations in BAP1 appear to be uniformly associated with poor prognosis in patients with RCC. Knowledge gaps remain with regard to the predictive relevance of BAP1 alterations, especially in the context of immunotherapy. Prospective studies are required to more precisely ascertain the predictive value of BAP1 alterations in RCC.
INTRODUCTION
BRCA1-associated protein 1 - BAP1 gene is a tumor suppressor located on the human chromosome 3p21.3 and encodes ubiquitin carboxy-terminal hydrolase. It is considered a classic two-hit tumor suppressor gene. [1] Within the nucleus, BAP1 acts as a chromatin scaffold for chromatin-remodeling complexes and hence regulates cell proliferation by deubiquitylating host cell factor 1 (HCF1). [2, 3] Cytoplasmic BAP1 is localized to the endoplasmic reticulum, where it stabilizes type 3 inositol-1,4,5-trisphosphate receptor (IP3R3), [3] which via calcium mediated cytochrome-c release from the mitochondria leads cell apoptosis. [1, 4]. Moreover, BAP1 has been shown to play a role in the metabolic activity of cells. For example, a study that evaluated plasma from sixteen BAP1 + /–individuals from 2 families carrying various germline BAP1 mutations and compared with thirty BAP1 wild-type (wt) controls from the same families [5]. They observed increased glycolysis and increased reduced aerobic mitochondrial respiration in BAP1 + /–, as compared to BAP1wt members, thus concluding that Warburg effect was seen in cells from individuals carrying heterozygous germline BAP1 mutations, much like cancer cells and these mutations may be the reason for a higher incidence of cancer among them [5].
Germline BAP1 mutations were observed in patients with familial mesotheliomas [6] and familial melanocytic tumors [7, 8]. A meta-analysis of all the published studies with BAP1-mutated families showed an increased association of BAP1 mutations with malignant mesothelioma, uveal melanoma and cutaneous melanomas, the etiology of a novel BAP1 cancer syndrome [8]. Analysis of clear-cell renal cell carcinoma (ccRCC), revealed the prevalence of BAP1mutations at 14% [9, 10]. Some studies showed correlation between loss of BAP1 activity and higher grade tumors [10, 11], and a molecular subtype of ccRCC with mutations in VHL and BAP1 was proposed. Since then, several studies have analyzed the role of BAP1 in clear-cell RCC prognosis as well as in defining responsiveness to various treatment modalities. However, here is paucity of data defining the role of BAP1 alterations in the context of response to immunotherapy in patients with ccRCC.
This systematic review was designed to evaluate the existing literature with regard to the clinical utility of BAP1 mutations in patients with metastatic ccRCC.
Diagnosis of BAP1 status
The use of immunohistochemistry (IHC) to identify BAP1 protein has previously been described in a cohort of 176 ccRCC tumor samples [10]. Of these, 148 had wild-type for BAP1 and 150 tumors showed the presence of nuclear BAP1 protein by IHC. Twenty-two samples carried the BAP1 mutation while 25 samples were negative for the BAP1 protein by IHC. This study showed that the positive and negative predictive values of the IHC for detection of BAP1 protein in ccRCC were > 98%. There is now a Clinical Laboratory Improvement Amendments (CLIA)–certified IHC test available for BAP1 protein available for use in clinical practice. More recently, non-invasive techniques such as radiomic features from CT scans are being evaluated to predict genomic status, including BAP1 mutations, in kidney tumors [12–14].
Variability in prevalence of BAP1 mutations
Unlike ccRCC, the role of BAP1 mutations is not as prominent in tumors of non-clear cell histology. In a cohort of patients with 186 ccRCC and 79 non-ccRCC, loss of BAP1 expression was seen in 9% (17/186) of the ccRCC tumors but only in 1% (1/79) of the non-ccRCC tumors (p = 0.016) [15]. Analysis of the TCGA dataset revealed prevalence of BAP1 mutations at 5.6% in papillary RCC (compared to 11% in ccRCC) [16]. Also BAP1 mutation correlated with decreased OS in the entire cohort (p = 0.0002) and within the ccRCC group (p = 0.0035); however, BAP1 mutation did not correlate with survival in papillary or chromophobe RCC. However, BAP1 mutations seem to have higher prevalence in patients with sarcomatoid RCC. In a cohort of 99 patients with sarcomatoid RCC and 906 patients with ccRCC, BAP1 mutations were found in 16% vs 9% respectively [17]. Recently, analysis of tumors from a total of 208 patients with sarcomatoid and rhabdoid RCC revealed significant enrichment for BAP1 somatic alterations when compared to classic ccRCC samples [18].
Sex/gender differences in BAP1 expression
There are differences in BAP1 expression between sexes and races. a higher prevalence of BAP1 mutations as reported in females vs. males and in black vs. white patients, however larger studies to validate these differences are required because results are variable between studies [19, 20]. A comprehensive analysis presented by Rickets and Linehan included a total of 628 sequenced samples (414 males and 214 females) with contributions from a US TCGA ccRCC cohort (424 total samples: 277 males and 147 females), a Japanese cohort (106 total samples: 78 males and 28 females) and a Chinese cohort (98 total sequenced samples: 59 males and 39 females) [21]. Of the total 10 genes analyzed, only BAP1 mutation rate was seen to be higher in tumors derived from female patients in a statistically significant manner (p = 0.0042). On the other hand, in a study analyzing 166 patients reported by Minardi et al, no significant correlation was observed between BAP1 expression and sex (p = 0.155) and age (p = 0.250) [22].
METHODS FOR SYSTEMATIC REVIEW
Evidence acquisition
Search strategy
A systematic literature search was conducted according to the PRISMA (Preferred Reporting Items for Systematic Reviews and Meta-Analyses) guidelines to identify studies reporting on BAP1 as a prognostic and/or predictive biomarker in RCC between 2010 and March 2021 [23]. The PubMed database was searched along with a free-text hand search using one or several combinations of the following items: BAP1, BRCA1-Associated Protein 1, ubiquitin carboxy-terminal hydrolase, ubiquitin thiolesterase AND clear cell renal cell carcinoma, ccRCC, conventional renal cell carcinoma, clear cell renal carcinoma, carcinoma, renal cell. The selection process was conducted in two stages; the first stage was used for initial screening of the title and abstract to identify eligible publications. The second stage was done via full-text reading including a manual search of publications in journals not listed in PubMed to further avoid missing any eligible study. For this systematic review, we excluded (I) non-English articles, (II) non-original articles (i.e., review articles with or without systematic review or meta-analysis), (III) editorials or case reports (IV) repeated publications on the same cohort to avoid publication bias.
Data extraction
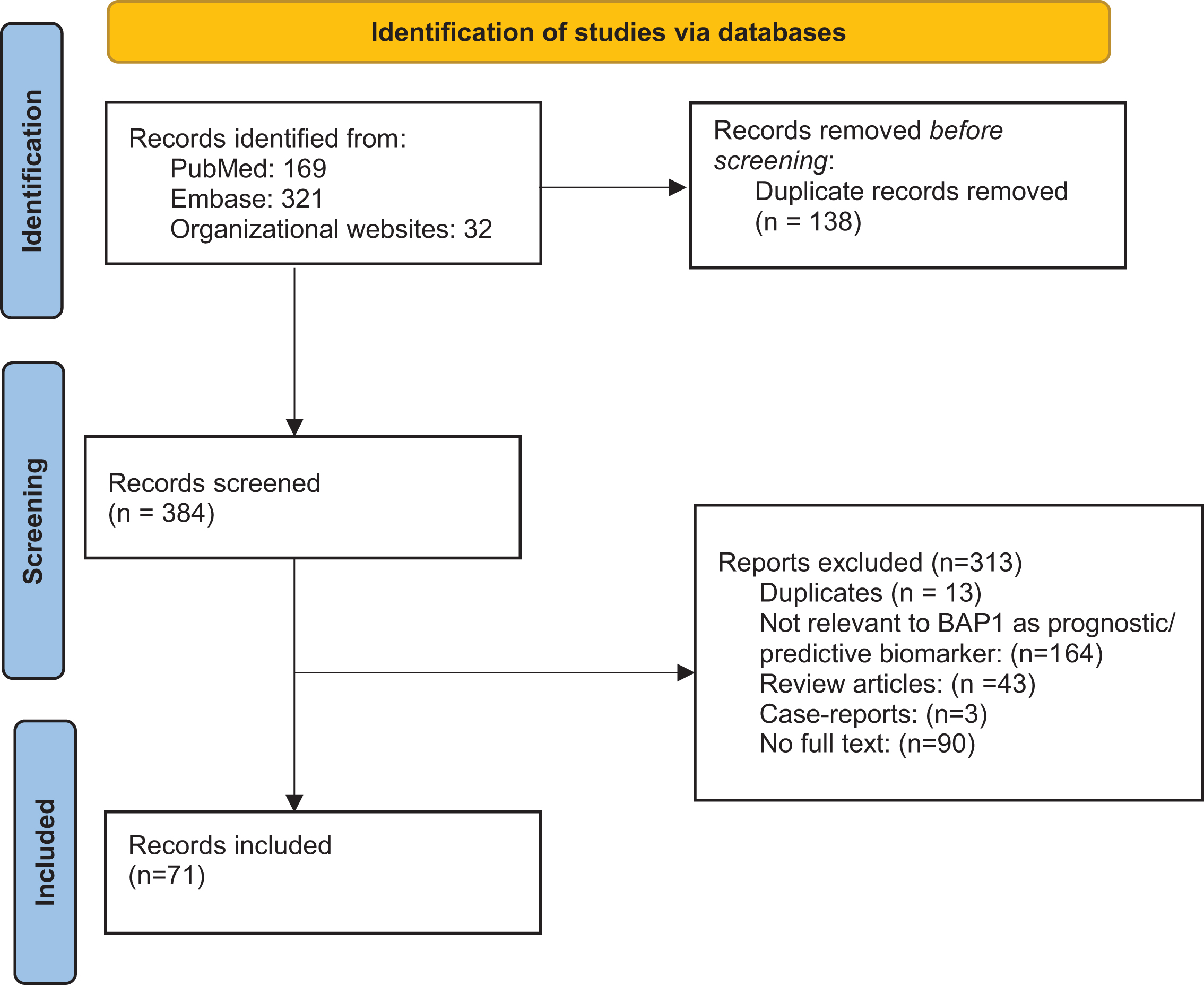
A CONSORT diagram for the selection process of included studies is provided in Fig. 1. The literature was searched for records focused on ccRCC and BAP1. Search strategies were created and run by a librarian using a combination of keywords and controlled vocabulary in the databases: PubMed and Embase.com. No filters or limits were applied to this search. ASCO, ASCO-GU, and ESMO conference proceedings were also searched using the same keywords. All search strategies were completed and run on April 25, 2021. Records were added to and deduplicated through EndNote and then uploaded and rechecked for duplicates in Rayyan. The final total was 384 unique records. After removal of abstracts without full text descriptions and removal of duplicates, we had a total of 71 that were then analyzed.
Fig. 1
PRISMA Flowchart.
RESULTS
Database search yielded a total of 522 citations, of which the title and abstract were screened for relevance. From these citations, 71 were subjected to full-text review, resulting in articles that met criteria for inclusion (Fig. 1).
Analysis was done in 2 categories. The first category includes studies where the role of BAP1 mutations is described as a prognostic biomarker by defining their association with tumor size, grade, stage at presentation, pathologic features of the tumor, and survival in localized as well as metastatic ccRCC (Table 1a and 1b). The second category (Table 2) includes studies where the role of BAP1 mutations is described as a predictive biomarker of responses to treatment regimens (TKI, mTOR inhibitors, nivolumab) in the metastatic setting.
Table 1a
Studies defining the role of BAP1 as a prognostic biomarker in localized ccRCC
| Article | Number of patients in analysis (N) | BAP1 prevalence (%) | Key findings implicating the prognostic role of BAP1 | Cancer outcomes |
| Samuel Peña-Llopis et al. [10] | N = 176 (76 discovery; 92 validation) | 14% | – BAP1 loss: correlated with high Fuhrman nuclear grade (q = 0.0005) | None reported |
| – tumors with loss of both BAP1 and PBRM1, had a significant association with rhabdoid features | ||||
| Sato et al. [38] (abstract) | N = 106 | 12% | – BAP1 mutations correlated with poor prognosis (features defining poor prognosis not defined) | None reported |
| Kapur et al. [24] | N = 472 UTSW cohort: 145 TCGA cohort: 327 | UTSW cohort: 14% TCGA cohort: 6% | – BAP1mutation: aggressive features in the tumor (higher grade, coagulative necrosis, advanced clinical stage) – BAP1 mutation associated with sarcomatoid/ rhabdoid histology | UTSW cohort: Worse OS in BAP 1 mutated vs. PBRM1 mutated (4.6 yrs vs. 10.6 yrs) TCGA cohort: Higher probability of death in BAP1 mutated cohort (HR 2.8; 95% CI: 1.4– 5.9; p = 0.004) |
| Hakimi et al. [39, 30] | N = 609 MSKCC cohort: 188 TCGA cohort: 421 | MSKCC cohort: 6.4% TCGA cohort: 9.7% | MSKCC cohort: – BAP1 mutated tumor presented with higher tumor stage, – BAP1 mutated tumor had higher Fuhrman nuclear grade (P = 0.03) | TCGA cohort: – Worse OS in BAP1 mutants vs wild type (31.2 mos (95% CI 23.2, NA) vs. 78.2 mos (95% CI 70.3, NA) – Worse cancer specific survival in BAP1 mutated (HR 7.71; 95% CI 2.08– 28.6; p = 0.002) |
| TCGA cohort: – BAP1 mutations associated with higher T stages (P = 0.004) | ||||
| – BAP1 mutation associated with higher nuclear grades (P = 0.02) | ||||
| – BAP1 mutation associated with larger tumor sizes (P = 0.002) | ||||
| – BAP1 mutation significantly associated with metastasis at presentation (P = 0.01) | ||||
| Gosage et al. | N = 128 | 14 (11%) | – BAP1 mutation significantly associated with metastasis at presentation (p = 0.037) – BAP1 mutation significantly associated with advanced clinical stage | – Shorter RFS in BAP1 mutated tumors compared to PBRM1 mutated tumors (75th centile for survival 1.22 years vs. 4.9 years; p = 0.059) |
| – No significant difference in OS in BAP1 mutant vs. PBRM1 mutated tumors | ||||
| Kapur et al. [40] (abstract) | N = 559 | 14.7% | BAP1 mutation associated with: – high Fuhrman grade (p < 0.0001) – advanced pT stage (p = 0.0021) – necrosis (p < 0.0001) – BAP1 mutation associated with sarcomatoid change (p = 0.0001) | – Worse DFS in BAP1 mutated vs. unmutated tumors (HR 2.9, 95% CI 1.8– 4.7, p < 0.0001) – Worse OS in BAP1 mutated vs. unmutated tumors (HR 2.0, 95% CI 1.3– 3.1, p = 0.0010) |
| Joseph et al. [41, 42] | N = 1,439 | 10.3% | BAP1 mutation associated significantly (all p < 0.0001): | – Higher probability of cancer related death (HR 3.06; 95% CI 2.28 – 4.10; p < 0.001) in BAP1 mutated tumors |
| – larger tumor size, | ||||
| – higher TNM stage | ||||
| – higher nuclear grade | ||||
| – coagulative tumor necrosis | ||||
| – higher SSIGN and UISS scores. | ||||
| Togo et al. | N = 45 | 11.1% | – Worse RFS in tumors with biallelic loss-of function BAP1 mutation | |
| (p = 0.046); difference did not persist after multivariate analysis when combined with age, T- stage, histological subtype and vascular invasion | ||||
| Minardi et al. [22] | N = 162 (Only described pT1 tumors) | Not reported | BAP1 staining≤10% associated with: | – DFS not significantly worse in BAP1 mutated tumors as compared to unmutated (log-rank test; p = 0.368) |
| – larger tumor size | ||||
| – presentation with metastasis (not significant) | ||||
| – Low nuclear BAP1 expression associated with higher tumor grade (p = 0.021) | ||||
| Wang et al. [44] | N = 26 ccRCC | 13% | None reported | None reported |
| Ricketts et al (TCGA). [16] | N = 488 ccRCC | 11% | – Shorter OS in BAP1 mutated tumors (p = 0.0035) in the ccRCC cohort (not in the papillary or chromophobe RCC cohorts) | |
| Da Costa et al. [45] | N = 441 (Included stage I/ II ccRCC only) | 24.3% | BAP1 expression significantly associated with: | – Worse DSS rate in BAP1 mutated tumors (84.1% vs. 95.8%; p < 0.001). |
| – higher pT stage (p < 0.0001) | – Worse RFS in BAP1 mutated tumors (72% vs. 95.2) | |||
| – larger tumor size (p < 0.0001) | ||||
| – higher ISUP grade (p < 0.0001) | ||||
| – lymphovascular invasion | ||||
| Oka et al. [25] | N = 35 (non-metastatic ccRCC with an IVC tumor thrombus) | 31.2% | None reported | – Worse median OS in BAP1 expressing vs. negative tumors (44.7 vs. 81.5 mos; p = 0.052) |
| – Worse median DFS in BAP1 expressing vs. negative tumors (10.0 vs. 26.0 mos; p = 0.011) (significance persisted on multi-variate analysis). | ||||
| Wi et al. [46] | N = 300 | 18.7% | BAP1 loss associated with high WHO/ ISUP grade (p = 0.002) | – No association seen between BAP1 expression and RFS and cancer-specific survival |
| Manley et al. [29] | N = 203 | 7.4% | None reported | – While worse OS was seen in BAP1 mutated tumors in unadjusted analysis; the difference became insignificant after adjusting for various characteristics (p = 0.100) |
| Schwen et al. (abstract) | N = 70 | 20% | BAP1 mutations associated with: | None reported |
| – high grade | ||||
| – upstaging to pT3a (p < 0.036) | ||||
| Park et al. [47] | N = 24; (6 had synchronous metastasis) | 25% | – Two-fold enrichment shown in BAP1 mutations (and KDM5C and FOXC2) in aggressive ccRCC | – Worse OS in BAP1 mutated tumors (p < 0.05) |
| Yang et al. [48] | N = 45 | 24% | BAP1 mutation significantly correlated with: | – Shorter OS in BAP1 mutated tumors (25.09±1.76 mos. vs. 31.91±2.02 mos; p < 0.05) |
| – larger tumor diameter | ||||
| – higher pathological stage | ||||
| – advanced TNM stage | ||||
| Gallan et al. [27] | N = 14 (ALL BAP1 mutated tumors) | BAP1 mutated tumors were significantly associated with: | None reported | |
| – higher stage (> T3) | ||||
| – renal vein invasion | ||||
| – 50% of BAP1 mutated tumors developed metastases | ||||
| Lin et al. [49] | N = 96 | 9% | BAP1 mutation associated with: | No difference in survival between BAP1 mutated vs. unmutated tumors. |
| – larger tumor size (P = 0.020) | ||||
| – higher tumor stage (P = 0.007) | ||||
| – higher rates of metastasis (P = 0.012) | ||||
| Bi et al. [19] | N = 105 | 9.5% | – BAP1 mutation (along with TP53 and PTEN) associated with higher grade and pTstage | None reported specifically for BAP1 mutated tumors. |
| – BAP1 (along with PTEN and ERBB2) associated with | ||||
| metastasis at diagnosis | ||||
| – BAP1 and MET associated with sarcomatoid differentiation |
Table 1b
Role of BAP1 mutations in metastatic RCC
| Article | Number of patients in analysis (N) | BAP1 prevalence (%) | Concordance between primary and metastatic sites for BAP1 mutation | Key findings implicating the prognostic role of BAP1; cancer related outcomes and conclusions |
| Shreders, A et al (abstract) | N = 99 ccRCC (48 M0 and 51 M1) | Not reported | 99% | None reported |
| Miura et al. [33] | N = 504 (103 patients developed recurrent RCC) | 19.5% at primary site 26.8% at metastatic site | – Concordance for BAP1: 63.4% | – Worse median OS for BAP1 mutated tumors (51 mos. vs. 97 mos; (P = 0.0077). |
| Becerra, M et al (abstract) [36] | N = 153 patients (ccRCC: 94) | 19.8% in ccRCC | Not reported | – Pleural metastases enriched for BAP1 mutations (p = 0.008) |
| Eckel-Passow et al. [32] | N = 97 ccRCC (M0 and M1) | 20% | – Concordance for BAP1: 98% (100% in metachronous and 96% in synchronous metastatic tumors) | – No statistically significantly association reported in CSS in metastatic lesions with BAP1 expression vs. not (HR = 1.29, 95% CI: 0.76– 2.19, p = 0.34) |
| Da Costa et al. [35] | N = 124 ccRCC (M1 = 124 38 paired cases from primary) | 62.1% metastatic lesions stained negative | – High discordance reported between BAP1 expression in primary vs metastatic tumor (44.7%) | – Worse OS rates in BAP1 negative tumors (35.1% vs. 53.2%; P = 0.004) – Worse PFS rates in BAP1 negative tumors (3.9% vs. 14.9%; P = 0.003) |
| Bossé et al. [50] (abstract) | N = 308 | 19% | Not reported | – Worse OS survival in BAP1 nutated tumors (aHR 1.7; 95% CI 1.1– 2.5, p = 0.01) |
| – BAP1 associated with worse IMDC risk group |
Table 2
Responsiveness to treatments in metastatic tumors
| Article | Number of patients in analysis (N) | BAP1 prevalence (%) | Drug evaluated in study | Correlation of BAP1 with response to specific drug |
| Lim et al. [56] | N = 15 ccRCC | Not reported | Everolimus | – BAP1 mutation seen in 2 patients with ccRCC and both lacked clinical benefit with everolimus. |
| Hsieh et al. [51] (RECORD-3 trial) | N = 220 mccRCC 1st line evero-limus = 109 1st line sunitinib = 111 | 19% | Compared 1st line everolimus followed by sunitinib at progression with the opposite sequence | – BAP1 mutated tumors showed shorter median PFS1L in both everolimus and sunitinib arms. |
| – BAP1 mutation also associated with worse OS compared to wild type tumors. | ||||
| Voss et al. [57] (COMPARZ trial) | N = 357 mRCC Sunitinib = 175 Pazopinib = 182 | 15% | Patients on trial received first-line sunitinib vs. pazopanib | – Worse OS in BAP1 mutated tumors regardless of treatment arm (log-rank, p = 0.012) |
| – On multivariate analysis, presence of BAP1 or TP53 mutations (either or both) and absence of PBRM1 mutation was independent drivers of worse outcomes. | ||||
| Carlo et al. | N = 105 mRCC | 24% | VEGF inhibitors | – Lower time to treatment failure in BAP1 mutated tumors vs. wild type (median 6.4 mos vs. 11.0 mos; p = 0.01) |
| – Shorter OS associated with BAP1 mutations | ||||
| García-Donas, J. et al. [52] (abstract) | N = 77 (87% with ccRCC) | Not reported | Everolimus (79%) vs. temsirolimus (21%) | – BAP1 mutated tumors associated with improved response to mTOR inhibitors |
| Voss et al. [58] (combined COMPARZ and RECORD-3 data) | COMPARZ N = 357 (training cohort) RECORD-3 N = 258 (validation cohort) | COMPARZ: 15% | TKI | – Lower OS in BAP1 mutated tumors (31.5 vs 22.1 months in wild type; p = 0·0261) |
| – updated MSKCC model with impact of mutations in 3 genes; validated in RECORD-3 cohort | ||||
| Ravaud et al. [59] Patients from S-TRAC phase-III trial (abstract) | N = 171 pts (Sunitinib treated N = 91) | Not reported | Sunitinib | – No impact of BAP1 mutation seen on DFS |
| Vano et al. [54] NIVOREN GETUG-AFU-26 trial (abstract) | N = 324 | Not reported | Nivolumab (PD-1 inhibitor) | – No impact of BAP1 mutations seen on OS. |
| Braun et al. [60] (combined data from CheckMate (CM) trials and TCGA)) | N = 454 patients from CM trials | 19% | Nivolumab Everolimus | – No impact of BAP1 mutation on response to drugs |
| [N = 261 treated with anti-PD-1] | ||||
| [N = 193 treated with mTORi] | ||||
| TCGA: N = 366 (20% metastatic tumors) |
Frequency and impact of BAP1 mutations on tumor characteristics and prognosis
We found an overall prevalence of BAP1 mutations in patients with non-metastatic early stage ccRCC between 6–24%. A review of all studies cited here uniformly depicted tumors with mutated BAP1 as carrying poor prognosis. Peña-Llopis et al were the first group to describe in a discovery cohort the correlation between BAP1 mutation and occurrence of high grade tumors [10]. Soon thereafter, Kapur et al combined 2 cohorts of almost 470 patients (including 327 tumors from the TCGA database) to describe in detail the correlation between BAP1 mutations and aggressive features on tumors including higher grade, sarcomatoid and rhabdoid features and coagulative tumor necrosis [24]. This was followed by several other studies that correlated BAP1 mutations with high-risk tumor characteristics as well as with adverse cancer related outcomes such as overall survival (OS), cancer specific survival, presence of metastatic disease at the time of diagnosis (Table 1). Studies identified a higher prevalence of BAP1 mutations, up to 31%, in patients that presented an IVC thrombus at the time of diagnosis, supporting the association with poor prognosis [25–27].
Studies have also looked specifically at patients with small renal masses (< 4 cm). A study that analyzed 70 samples from T1 tumors (of which 20% had BAP1 mutated tumors) found a significant association between BAP1 and high grade tumors [28]. Another cohort of 203 small renal tumors, found a correlation between BAP1 mutations and poor survival in unadjusted analysis (P = 0.050), however the difference became insignificant after adjustment for multiple factors (adjusted P = 0.100) [29].
As further studies continued to evolve, the absence of BAP1 protein has also been correlated with early metastasis in patients that were followed after initial nephrectomies [19, 22, 27, 30, 31]. Studies describing an evaluation of primary and metastatic lesions in patients with ccRCC for BAP1 mutations are shown in Table 1b. Overall, > 80% concordance was found between the primary and metastatic sites for BAP1 mutations in most studies [32–34]. One study by daCosta et al showed ∼45% discordance between BAP1 in primary vs metastatic tumor sites [35]. Interestingly, metastatic lesions to the pleura were enriched for BAP1 mutations [36] while these mutations were infrequent in patients with pancreatic metastasis, supporting an indolent course for the latter [37].
BAP1 as a predictive biomarker
Management of kidney cancer has undergone a paradigm shift with the approval of many new therapies over the last two decades. However, we have not yet been able to identify molecular targets to predict response to specific therapies. The role of BAP1 mutation as a predictor of responsiveness to targeted agents has been described from analysis of the RECORD-3 and COMPARZ phase-III clinical trials. Previous retrospective studies found an association between BAP1 mutations and mTOR pathway activation [24, 10]. No such data are currently available from prospective immunotherapy trials and thus there remains an existing knowledge gap.
i) Response to mTOR inhibitors: A study conducted by Lim et al. that included several cancers treated with the mTOR inhibitor, everolimus, included 15 patients with metastatic RCC; mutated BAP1 was noted only in 2 patients without a response to everolimus. The results from RECORD-3, a phase -III study comparing first-line everolimus followed by sunitinib (VEGF- TKI) at progression with the opposite sequence in patients who experienced progression [51]. For everolimus-treated patients, those with BAP1 mutated cancers were seen to have a higher risk of progression than those with wild-type BAP1 (median PFS first line (IL) of 4.9 vs 10.5 months; hazard ratio (HR): 1.84; 95% CI: 1.1, 3.2). Similarly, in the sunitinib arm tumors with mutated BAP1 had a higher risk of progression than tumors with wild-type BAP1 (median PFS 1L 8.1 vs 11.0 months); the HR here was not significant (HR: 1.69; 95% CI: 0.9, 3.2). This indicated that cancers with BAP1 mutants had a poor prognosis regardless of treatment regimen (mTOR inhibitor vs. TKI) used in the front-line setting. Contradictory to these two studies, the Spanish Oncology Genitourinary Group (SOGUG) presented data on 77 patients with kidney cancer (of whom 87% had clear cell RCC). In these everolimus- (79%) or temsirolimus-(21%) treated patients, lack of IHC expression for BAP1 was associated with better mTOR inhibitor response, even on multivariable analysis [52].
ii) Response to VEGF-TKIs: As stated above, in the RECORD-3 clinical trial, even though not significant, sunitinib treated patients had a higher risk of progression in the presence of a BAP1 mutations as compared to wild type BAP1 (median PFS 1L 8.1 vs 11.0 months; HR: 1.69; 95% CI: 0.9, 3.2). A retrospective study from an institutional cohort of patients at MSKCC (n = 105) that included 24% patients with BAP1 mutations showed a shorter time to treatment failure for patients with mutated BAP1 in response to VEGF-TKIs (median 6.4 months vs 11.0 months; p = 0.01) as well as a shorter overall survival (median 28.7 months vs. not reached; p = 0.02) [53].
iii) Response to immunotherapy-based therapy: Even though data from larger clinical trials using immune checkpoint inhibitors is lacking, smaller studies have tried to dissect this relationship. The NIVOREN GETUG-AFU 26 study included 324 patients who had received the programmed death-1 (PD-1) inhibitor nivolumab [54]. The investigators reported no association of BAP-1 loss with PFS or OS (p = 0.6 and 0.9 respectively). Braun et al described 592 patients on clinical trials- CheckMate 010/009 (phase-II) and CheckMate 025 (a phase III trial that demonstrated an OS benefit with nivolumab over the mTOR inhibitor everolimus in previously treated patients with ccRCC) [55]. The study included 261 patients treated with PD-1 inhibitor and 193 patients treated with mTOR inhibition along with predominantly localized ccRCC tumors from the TCGA dataset. Overall, they reported prevalence of BAP1 mutation at around 19% in advanced ccRCC and no association was reported between BAP1 mutation and response to nivolumab or mTOR inhibition.
DISCUSSION
In this systematic review, we summarize the studies investigating prognostic and predictive role of BAP1 mutations in RCC. In most studies included in this review, BAP1 alterations portended a worse prognosis as compared to the patients without these alterations. In most of the studies, BAP1 mutations correlate with tumor characteristics such as higher-grade, presence of necrosis, larger size, and sarcomatoid/ rhabdoid change. Importantly, tumors with mutated BAP1 seem to have worse prognosis regardless of treatment regimen, although controversies exist between trials. In large phase-III clinical trials, COMPARZ and RECORD-3; BAP1 mutations led to poor outcomes when treated with VEGF-inhibitors. However, a similar analysis from the phase-III S-TRAC trial (using adjuvant sunitinib for patients with stage III ccRCC) did not show an impact of BAP1 alterations on DFS. Only small studies as shown in Table 2, have tried to correlate these mutations with outcomes when treated with immune checkpoint inhibitor and have not found an association. Previous studies have, however, shown BAP1 mutations as potentially related to markers of responsiveness to immune checkpoint inhibitors. For example, BAP1mutation prevalence has been seen to be higher in sarcomatoid and rhabdoid tumors and these tumors are known to be more responsive to immunotherapy drugs [18]. Another study by Pal et al on samples from 648 patients has shown the average tumor mutation burden to be higher in ccRCC samples with co-occurring BAP1 and PBRM1 mutations [61]. Further studies have shown an association between an inflammatory tumor microenvironment at BAP1 loss as well [62]. Wang et al. identified an “inflamed” subtype of RCC, which was enriched for BAP1 mutations while the “non-inflamed” subtype was enriched for angiogenesis-related genes. Similar findings were reported from a real-world patient population of 316 patients, where a higher prevalence of BAP1 mutations was found in the “inflamed” or T-effector subgroup (18.6% vs. 3.0%, p = 0.0035) [63]. The role of BAP1 alterations non- clear cell RCC is much less studies due to the lower prevalence of these tumors as compared to the clear-cell type.
This systematic review has several limitations. While performing the search, it was evident that there were not enough studies that reported numerical data on survival and response to treatment in the context of BAP1 mutations. Due to paucity of available data, a merged analysis of outcomes was not possible. Moreover, while we meticulously tried to focus on metastatic RCC and their responsiveness to immune checkpoint inhibitors, some studies presented included non-metastatic patients as well. Thus, the role of BAP1 as a predictive marker based on the available data could not be optimally defined.
Further studies to evaluate the role of BAP1 alterations as part of a broader need to find the optimal biomarker in RCC are required. As a variety of effective therapies become available and more trials are ongoing for patients with kidney cancer, it is imperative to discover better predictive and prognostic biomarkers. This has eluded RCC so far. It seems that a single biomarker such as a single gene mutation or a single gene expression signature will likely not be helpful in predicting risk of recurrence or response to treatments. Instead, it is imperative to study various biomarkers as part of a “composite predictive and prognostic biomarker” in ccRCC [64]. We recommend integrating a stratifying approach in forthcoming clinical trials for localized as well as advanced RCC, where a multidimensional integrated biomarker is incorporated; such that it involves tumor genomic features, including mutations such as BAP-1, transcriptomic profiles and other biomarkers of interest such as those related to response to various therapies.
ACKNOWLEDGMENTS
The authors would like to acknowledge support as listed below:
SG: The project described was supported by the National Center for Advancing Translational Sciences of the National Institutes of Health, under Award Number 2KL2TR001426-05A1. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH.
PNL: Dr. Lara is supported in part by the generosity of Gerry and Susan Knapp and by NCI Cancer Center Support Grant P30 CA093373.
FUNDING
The authors report no funding received for this work.
AUTHOR CONTRIBUTIONS
SG, PNL: conception, design, interpretation of results, manuscript review and revision. MP helped with extraction of studies. SG conducted the statistical analysis and drafted the manuscript.
CONFLICT OF INTEREST
SG reports no relevant conflicts of interest
MP reports no relevant conflicts of interest
PNL reports no relevant conflicts of interest
PNL is an Editor-in-Chief of this journal, but was not involved in the peer-review process of this paper, nor had access to any information regarding its peer-review.
REFERENCES
[1] | Bononi A , Giorgi C , Patergnani S , et al. BAP1 regulates IP3R3-mediated Ca 2+ flux to mitochondria suppressing cell transformation. Nature. (2017) ;546: (7659):549–553. doi: 10.1038/nature22798 |
[2] | BrugarolasJ. PBRM1 and BAP1 as novel targets for renal cell carcinoma. Cancer J. (2013) ;19: (4):324–332. doi:10.1097/PPO.0b013e3182a102d1 |
[3] | Carbone M , Harbour JW , Brugarolas J , et al. Biological Mechanisms and Clinical Significance of BAP1 Mutations in Human Cancer. Cancer Discov. (2020) ;10: (8):1103–1120. doi: 10.1158/2159-8290.CD-19-1220 |
[4] | Zhang Y , Shi J , Liu X , et al. BAP1 links metabolic regulation of ferroptosis to tumour suppression. Nature Cell Biology. (2018) ;20: (10):1181–1192. doi: 10.1038/s41556-018-0178-0 |
[5] | Bononi A , Yang H , Giorgi C , et al. Germline BAP1 mutations induce a Warburg effect. Cell Death & Differentiation. (2017) ;24: (10):1694–1704. doi: 10.1038/cdd.2017.95 |
[6] | Testa JR , Cheung M , Pei J , et al. Germline BAP1 mutations predispose to malignant mesothelioma. Nat Genet. (2011) ;43: (10):1022–1025. doi: 10.1038/ng.912 |
[7] | Wiesner T , Obenauf AC , Murali R , et al. Germline mutations in BAP1 predispose to melanocytic tumors. Nat Genet. (2011) ;43: (10):1018–1021. doi: 10.1038/ng.910 |
[8] | Carbone M , Ferris LK , Baumann F , et al. BAP1 cancer syndrome: malignant mesothelioma, uveal and cutaneous melanoma, and MBAITs. Journal of Translational Medicine. (2012) ;10: (1):179. doi: 10.1186/1479-5876-10-179 |
[9] | Guo G , Gui Y , Gao S , et al. Frequent mutations of genes encoding ubiquitin-mediated proteolysis pathway components in clear cell renal cell carcinoma. Nature Genetics. (2012) ;44: (1):17–19. doi: 10.1038/ng.1014 |
[10] | Peña-Llopis S , Vega-Rubín-de-Celis S , Liao A , et al. BAP1loss defines a new class of renal cell carcinoma. Nature Genetics. (2012) ;44: (7):751–759. doi: 10.1038/ng.2323 |
[11] | Turajlic S , Xu H , Litchfield K , et al. Tracking Cancer Evolution Reveals Constrained Routes to Metastases: TRACERx Renal. Cell. (2018) ;173: (3):581–594.e12. doi: 10.1016/j.cell.2018.03.057 |
[12] | Zeng H , Chen L , Wang M , Luo Y , Huang Y , Ma X . Integrative radiogenomics analysis for predicting molecular features and survival in clear cell renal cell carcinoma. Aging (Albany NY). (2021) ;13: (7):9960–9975. doi: 10.18632/aging.202752 |
[13] | Chen X , Zhou Z , Hannan R , et al. Reliable gene mutation prediction in clear cell renal cell carcinoma through multi-classifier multi-objective radiogenomics model. Phys Med Biol. (2018) ;63: (21):215008. doi: 10.1088/1361-6560/aae5cd |
[14] | Meza LA , Filippov A , Naim S , et al. Radiomic features of renal cell carcinoma primary and metastatic sites as predictors of TERT and BAP1 mutations. JCO. (2021) ;39: (6_suppl):282–282. doi: 10.1200/JCO.2021.39.6_suppl.282 |
[15] | Ho TH , Kapur P , Joseph RW , et al. Loss of PBRM1 and BAP1 Expression Is Less Common in Non–Clear Cell Renal Cell Carcinoma Than in Clear Cell Renal Cell Carcinoma. Urol Oncol. (2015) ;33: (1):23.e9–23.e14. doi: 10.1016/j.urolonc.2014.10.014 |
[16] | Ricketts CJ , De Cubas AA , Fan H , et al. The Cancer Genome Atlas Comprehensive Molecular Characterization of Renal Cell Carcinoma. Cell Reports. (2018) ;23: (1):313–326.e5. doi: 10.1016/j.celrep.2018.03.075 |
[17] | Casuscelli Jozefina , Manley Brandon J , Mano Roy , et al. Mp71-14 analysis of the mutational landscape of sarcomatoid clear cell renal cell carcinoma compared to conventional clear cell renal cell carcinoma. Journal of Urology. (2016) ;195: (4S):e921–e921. doi: 10.1016/j.juro.2016.02.1460 |
[18] | Bakouny Z , Braun DA , Shukla SA , et al. Integrative molecular characterization of sarcomatoid and rhabdoid renal cell carcinoma. Nature Communications. (2021) ;12: (1):808. doi: 10.1038/s41467-021-21068-9 |
[19] | Bi H , Yin J , Zhou L , et al. Clinicopathological and prognostic impact of somatic mutations in Chinese patients with clear cell renal cell carcinoma. Translational Andrology and Urology. (2752) ;9: (6):763–763. doi: 10.21037/tau-20-1410 |
[20] | Paulucci DJ , Sfakianos JP , Yadav SS , Badani KK . BAP1 is overexpressed in black compared with white patients with Mx-M1 clear cell renal cell carcinoma: A report from the cancer genome atlas. Urol Oncol. (2016) ;34: (6):259.e9–259.e14. doi: 10.1016/j.urolonc.2015.12.019 |
[21] | Ricketts CJ , Linehan WM . Gender Specific Mutation Incidence and Survival Associations in Clear Cell Renal Cell Carcinoma (CCRCC). PLOS ONE. (2015) ;10: (10):e0140257. doi: 10.1371/journal.pone.0140257 |
[22] | Minardi D , Lucarini G , Milanese G , Montironi R , Di Primio R . Prognostic role of BAP1 in pT1 clear cell carcinoma in partial nephrectomy specimens. Virchows Arch. (2017) ;471: (1):99–105. doi: 10.1007/s00428-017-2143-x |
[23] | PRISMA. Accessed June 14, 2021. http://prisma-statement.org/prismastatement/flowdiagram.aspx |
[24] | Kapur P , Peña-Llopis S , Christie A , et al. Effects on survivalof BAP1 and PBRM1 mutations in sporadic clear-cell renal cellcarcinoma: a retrospective analysis with independent validation. Lancet Oncol. (2013) ;14: (2):159–167. doi: 10.1016/S1470-2045(12)70584-3 |
[25] | Oka S , Inoshita N , Miura Y , et al. The loss of BAP1 protein expression predicts poor prognosis in patients with nonmetastatic clear cell renal cell carcinoma with inferior vena cava tumor thrombosis. Urologic Oncology: Seminars and Original Investigations. (2018) ;36: (8):365.e9–365.e14. doi: 10.1016/j.urolonc.2018.04.015 |
[26] | Liu C , Gong X , Zhang S , et al. PT321 - Comprehensive molecular characterization of clear cell renal cell carcinoma with caval tumour thrombus. European Urology Supplements. (2019) ;18: (1):e2100. doi: 10.1016/S1569-9056(19)31522-2 |
[27] | Gallan AJ , Parilla M , Segal J , Ritterhouse L , Antic T . BAP1-Mutated Clear Cell Renal Cell Carcinoma: A Clinicopathologic Characterization of 14 Molecularly Confirmed Tumors. American Journal of Clinical Pathology. (2021) ;155: (5):718–728. doi: 10.1093/ajcp/aqaa176 |
[28] | MP08-07 GENOMIC PROFILING OF CT1A CLEAR CELL RENAL CELL CARCINOMA FOR PREDICTING AGGRESSIVE PATHOLOGY | Journal of Urology. Accessed May 25, 2021. https://www.auajournals.org/doi/10.1097/JU.0000000000000828.07 |
[29] | Manley BJ , Reznik E , Ghanaat M , et al. Characterizing recurrent and lethal small renal masses in clear cell renal cell carcinoma using recurrent somatic mutations. Urologic Oncology: Seminars and Original Investigations. (2019) ;37: (1):12–17. doi: 10.1016/j.urolonc.2017.10.012 |
[30] | Hakimi AA , Chen Y-B , Wren J , et al. Clinical and pathologic impact of select chromatin-modulating tumor suppressors in clear cell renal cell carcinoma. Eur Urol. (2013) ;63: (5):848–854. doi: 10.1016/j.eururo.2012.09.005 |
[31] | Clinical and pathological impact of VHL, PBRM1, BAP1, SETD2,KDM6A,and JARID1c in clear cell renal cell carcinoma –Gossage –2014 –Genes, Chromosomes and Cancer - Wiley Online Library. Accessed May 18, 2021. https://onlinelibrary.wiley.com/doi/full/10.1002/gcc.22116 |
[32] | Eckel-Passow JE , Serie DJ , Cheville JC , et al. BAP1 and PBRM1 in metastatic clear cell renal cell carcinoma: tumor heterogeneity and concordance with paired primary tumor. BMC Urol. (2017) ;17: . doi: 10.1186/s12894-017-0209-3 |
[33] | Miura Y , Inoshita N , Ikeda M , et al. Loss of BAP1 protein expression in the first metastatic site predicts prognosis in patients with clear cell renal cell carcinoma. Urol Oncol. (2017) ;35: (6):386–391. doi: 10.1016/j.urolonc.2017.02.003 |
[34] | Shreders A , Joseph RW , Serie D , et al. High concordance of BAP1 and PBRM1 expression in patient-matched primary and metastatic ccRCC tumors. JCO. (2015) ;33: (7_suppl):507–507. doi: 10.1200/jco.2015.33.7_suppl.507 |
[35] | da Costa WH , Fares AF , Bezerra SM , et al. Loss of BAP1 expression in metastatic tumor tissue is an event of poor prognosis in patients with metastatic clear cell renal cell carcinoma. Urologic Oncology: Seminars and Original Investigations. (2019) ;37: (1):78–85. doi: 10.1016/j.urolonc.2018.10.017 |
[36] | Abstracts. BJU International. (2016) ;118: (S5):4–30. doi: https://doi.org/10.1111/bju.13694 |
[37] | Singla N , Onabolu O , Woolford L , et al. Unraveling the molecular profile underpinning pancreatic tropisms in metastatic clear cell renal cell carcinoma. JCO. (2019) ;37: (15_suppl):e16096–e16096. doi: 10.1200/JCO.2019.37.15_suppl.e16096 |
[38] | Abstract 3184: Integrative analysis of clear cell renal cell carcinoma. | Cancer Research. Accessed May 18, 2021. https://cancerres.aacrjournals.org/content/73/8_Supplement/3184 |
[39] | Adverse Outcomes in Clear Cell Renal Cell Carcinoma with Mutations of 3p21 Epigenetic Regulators BAP1 and SETD2: A Report by MSKCC and the KIRC TCGA Research Network | Clinical Cancer Research. Accessed May 18, 2021. https://clincancerres.aacrjournals.org/content/19/12/3259.long |
[40] | Kapur Payal , Christie Alana , Raman Jay D , et al. BAP1 Immunohistochemistry Predicts Outcomes in a Multi-Institutional Cohort with Clear Cell Renal Cell Carcinoma. Journal of Urology. (2014) ;191: (3):603–610. doi: 10.1016/j.juro.2013.09.041 |
[41] | Joseph RW , Kapur P , Serie DJ , et al. Loss of BAP1 Protein Expression Is an Independent Marker of Poor Prognosis in patients with Low Risk Clear Cell Renal Cell Carcinoma. Cancer. (2014) ;120: (7):1059–1067. doi: 10.1002/cncr.28521 |
[42] | Joseph RW , Kapur P , Serie DJ , et al. Clear Cell Renal Cell Carcinoma Subtypes Identified by BAP1 and PBRM1 Expression. J Urol. (2016) ;195: (1):180–187. doi: 10.1016/j.juro.2015.07.113 |
[43] | Togo Y , Yoshikawa Y , Suzuki T , et al. Genomic profiling of the genes on chromosome 3p in sporadic clear cell renal cell carcinoma. International Journal of Oncology. (2016) ;48: (4):1571–1580. doi: 10.3892/ijo.2016.3395 |
[44] | Wang J , Xi Z , Xi J , et al. Somatic mutations in renal cell carcinomas from Chinese patients revealed by whole exome sequencing. Cancer Cell Int. (2018) ;18: . doi: 10.1186/s12935-018-0661-5 |
[45] | da Costa WH , da Cunha IW , Fares AF , et al. Prognostic impact of concomitant loss of PBRM1 and BAP1 protein expression in early stages of clear cell renal cell carcinoma. Urologic Oncology: Seminars and Original Investigations. (2018) ;36: (5):243.e1–243.e8. doi: 10.1016/j.urolonc.2018.01.002 |
[46] | Wi YC , Moon A , Jung MJ , et al. Loss of Nuclear BAP1 Expression Is Associated with High WHO/ISUP Grade in Clear Cell Renal Cell Carcinoma. J Pathol Transl Med. (2018) ;52: (6):378–385. doi: 10.4132/jptm.2018.09.21 |
[47] | Park JS , Pierorazio PM , Lee JH , et al. Gene Expression Analysis of Aggressive Clinical T1 Stage Clear Cell Renal Cell Carcinoma for Identifying Potential Diagnostic and Prognostic Biomarkers. Cancers. (2020) ;12: (1):222. doi: 10.3390/cancers12010222 |
[48] | Yang J , Gong C , Zhao J , et al. Effects of BAP1, Ki-67 index, and Id-1 in patients with clear cell renal carcinoma and their correlation with clinical features and prognosis. Translational Andrology and Urology. (2242) ;9: (5):250–250. doi: 10.21037/tau-20-1258 |
[49] | Lin P-H , Huang C-Y , Yu K-J , et al. Genomic characterization of clear cell renal cell carcinoma using targeted gene sequencing. Oncology Letters. (2021) ;21: (2):1–1. doi: 10.3892/ol.2021.12430 |
[50] | Bossé D , Xie W , Ged Y , et al. Alterations in key clear cell renal cell carcinoma (RCC) genes to refine patient prognosis. JCO. (2018) ;36: (15_suppl):4516–4516. doi: 10.1200/JCO.2018.36.15_suppl.4516 |
[51] | Hsieh JJ , Chen D , Wang PI , et al. Genomic Biomarkers of a Randomized Trial Comparing First-line Everolimus and Sunitinib in Patients with Metastatic Renal Cell Carcinoma. Eur Urol. (2017) ;71: (3):405–414. doi: 10.1016/j.eururo.2016.10.007 |
[52] | GarcÃa-Donas J , Roldan JM , Lainez N , et al. Comprehensivemolecular and immunohistochemical analysis of advanced renal cellcarcinoma patients treated with mTOR inhibitors. JCO. (2018) ;36: (15_suppl):4559–4559. doi: 10.1200/JCO.2018.36.15_suppl.4559 |
[53] | Carlo MI , Manley B , Patil S , et al. Genomic Alterations and Outcomes with VEGF-Targeted Therapy in Patients with Clear Cell Renal Cell Carcinoma. Kidney Cancer 1: (1):49–56. doi: 10.3233/KCA-160003 |
[54] | Vano Y-A , Rioux-Leclercq N , Dalban C , et al. NIVOREN GETUG-AFU 26 translational study: Association of PD-1, AXL, and PBRM-1 with outcomes in patients (pts) with metastatic clear cell renal cell carcinoma (mccRCC) treated with nivolumab (N). JCO. (2020) ;38: (6_suppl):618–618. doi: 10.1200/JCO.2020.38.6_suppl.618 |
[55] | Motzer RJ , Escudier B , McDermott DF , et al. Nivolumab versus Everolimus in Advanced Renal Cell Carcinoma. N Engl J Med. (2015) ;373: (19):1803–1813. doi: 10.1056/NEJMoa1510665 |
[56] | Lim SM , Park HS , Kim S , et al. Next-generation sequencing reveals somatic mutations that confer exceptional response to everolimus. Oncotarget. (2016) ;7: (9):10547–10556. doi: 10.18632/oncotarget.7234 |
[57] | Abstracts from the Sixteenth International Kidney Cancer Symposium, 2nd-3rd November 2017, Miami, Florida - IOS Press. Accessed May 18, 2021. https://content.iospress.com/articles/kidney-cancer/kca189001 |
[58] | Voss MH , Reising A , Cheng Y , et al. Genomically annotated risk model for advanced renal-cell carcinoma: a retrospective cohort study. The Lancet Oncology. (2018) ;19: (12):1688–1698. doi: 10.1016/S1470-2045(18)30648-X |
[59] | Ravaud A , Martini J-F , Ching K , et al. Phase III trial of adjuvant sunitinib in patients with high-risk renal cell carcinoma: Comprehensive tumour genomic and transcriptomic analyses. Annals of Oncology. (2019) ;30: :v358–v359. doi: 10.1093/annonc/mdz249.005 |
[60] | Braun DA , Hou Y , Bakouny Z , et al. Interplay of somatic alterations and immune infiltration modulates response to PD-1 blockade in advanced clear cell renal cell carcinoma. Nature Medicine. (2020) ;26: (6):909–918. doi: 10.1038/s41591-020-0839-y |
[61] | Pal SK , Madison R , Chung J , et al. Comparison of tumor mutational burden (TMB) in PBRM1/BAP1-based subsets of advanced renal cell carcinoma (aRCC). JCO. (2018) ;36: (6_suppl):634–634. doi: 10.1200/JCO.2018.36.6_suppl.634 |
[62] | Wang T , Lu R , Kapur P , et al. An Empirical Approach Leveraging Tumorgrafts to Dissect the Tumor Microenvironment in Renal Cell Carcinoma Identifies Missing Link to Prognostic Inflammatory Factors. Cancer Discov. (2018) ;8: (9):1142–1155. doi: 10.1158/2159-8290.CD-17-1246 |
[63] | Barata PC , Gulati S , Elliott A , et al. Angiogenic and T-effector subgroups identified by gene expression profiling (GEP) and propensity for PBRM1 and BAP1 alterations in clear cell renal cell carcinoma (ccRCC). JCO. (2021) ;39: (6_suppl):343–343. doi: 10.1200/JCO.2021.39.6_suppl.343 |
[64] | Gulati S , Vogelzang NJ . Biomarkers in renal cell carcinoma: Arewethere yet. Asian Journal of Urology. Published online June 6, 2021. doi: 10.1016/j.ajur.2021.05.013 |




