Association of SIRT1 single gene nucleotide polymorphisms and serum SIRT1 levels with laryngeal squamous cell carcinoma patient survival rate
Abstract
BACKGROUND:
SIRT1 is a multifunctional protein, possibly essential in tumorigenesis pathways, which can act both as a tumor promoter and tumor suppressor depending on the oncogenes, specific to particular tumors. Pathogenesis of laryngeal cancer is multifactorial and the association of SIRT1 expression with the clinical characteristics and prognosis of LSCC has not been fully identified.
OBJECTIVES:
The study aimed to evaluate associations between single gene nucleotide polymorphisms (SNPs) of SIRT1 (rs3818292, rs3758391, and rs7895833), serum SIRT1 levels, and 5-year survival rate in patients with laryngeal squamous cell carcinoma (LSCC).
METHODS:
The study involved 302 patients with LSCC and 409 healthy control subjects. The genotyping of SNPs was performed using RT-PCR, and serum SIRT1 levels were determined by the ELISA method.
RESULTS:
Our study found significant differences in genotype distributions of SIRT1 rs3758391 polymorphisms between the study groups. SIRT1 rs3758391 T/T genotype was associated with the increased LSCC development odds (OR
CONCLUSION:
The present study indicated a statistically significant association between the SIRT1 rs3758391 T/T genotype and increased LSCC development odds. LSCC patients with SIRT1 rs3818292 AG genotype showed a tendency to manifest with lower SIRT1 serum levels. No associations between SIRT1 SNPs and the 5-year survival rate of LSCC patients were detected.
1.Introduction
Laryngeal squamous cell carcinoma (LSCC) is one of the most common malignancies of the upper aerodigestive tract, with high mortality and poor prognosis for patients. In 2020, 184615 new cases of LSCC were diagnosed, and 99840 patients died from this cancer worldwide [1]. World Health Organization has estimated that the incidence and mortality of LSCC will have increased from 52.4% to 64.5%, respectively, by 2040 [2]. The most significant increment is projected in countries with a low Human Development Index [2]. This cancer occurs much more frequently in men than in women. According to The Global Cancer Observatory, the age-standardized incidence rate of laryngeal cancer is 3.6/100,000 people among males and 0.49/1,00,000 people among females per year [1]. This can be explained by the fact that men are more likely to have harmful habits. Laryngeal cancer is a multifactorial disease; however, it is generally accepted that tobacco and high-intensity alcohol consumption are the main risk factors for LSCC [3, 4, 6, 7]. Epidemiological studies have consistently linked pro-inflammatory dietary intake, human papillomavirus infection, environmental and occupational exposures with increased risk of LSCC [8, 9, 10, 11, 12, 13, 14]. A rising focus on identifying genetic and epigenetic factors to develop new diagnostic tools and treatment for laryngeal cancer can be seen [15]. Several studies have shown an essential role of gene variants, especially in combination with toxins, such as alcohol and cigarette smoke, in the etiopathology and the overall risk/survival rate of LSCC [15].
Despite the rather evident symptoms (hoarseness, dysphonia, dyspnea, and swallowing dysfunction) that may onset quite early in patients with laryngeal malignancies, in many cases, cancer already is at the advanced stage (III or IV) of the disease when diagnosed. Unfortunately, modern treatment strategies do not achieve improvement in therapeutic outcomes and survival [16, 17]. Therefore, to implement effective strategies for the prevention and early diagnosis, LSCC needs to be determined from epidemiological and clinical data and modern diagnostic methods based on genetic discoveries.
A silent information regulator two homolog 1 (SIRT1) is a nicotinamide adenine dinucleotide (NAD
Genetic variants may cause different SIRT1 expressions, and it could play an essential role in laryngeal carcinogenesis. However, just one study has investigated the involvement of SIRT1 single nucleotide polymorphism (SNP) in laryngeal cancer and has not found any associations between LSCC and SIRT1 rs12778366 gene polymorphism [34].
The study aimed to evaluate associations between SNPs of SIRT1 (rs3818292, rs3758391, and rs7895833), serum SIRT1 levels, and the 5-year survival rate of patients with LSCC.
2.Materials and methods
2.1Ethics statement
This case-control study was conducted at the Department of Otorhinolaryngology, Lithuanian University of Health Sciences (LUHS), Kaunas, Lithuania. The research protocol was approved by the Kaunas Regional Ethics Committee for Biomedical Research (authorization number BE-2-37). All procedures performed in the study followed the institution’s ethical standards, the Helsinki declaration, and its later amendments or similar ethical standards. Informed consent was obtained from all individual participants included in the study.
2.2Study protocol/design
Study population. A total study group consisted of 711 subjects: 302 patients with LSCC and 409 healthy individuals as a control group. The LSCC patient group included 292 (96.7%) males and 10 (3.3%) females. The control group included 393 (96.1%) males and 16 (3.9%) females. The LSCC patient and control groups were adjusted by sex and age. Demographic characteristics of the study groups are presented in Table 1.
Table 1
Demographic characteristics of the study
| Characteristic | Group | ||
|---|---|---|---|
| LSCC | Control group | ||
| Male, | 292 (96.7) | 393 (96.1) | 0.673* |
| Female | 10 (3.3) | 16 (3.9) | |
| Age years; mean (SD) | 63.2 (8.1) | 63.3 (11.6) | 0.146** |
| Smoking ***, | |||
| Yes | 124 (41.1) | 33 (8.1) | |
| No | 4 (1.3) | 156 (38.1) | |
| Alcohol consum- ption***, | |||
| Yes | 101 (33.4) | 102 (24.9) | |
| No | 27 (8.9) | 87 (21.3) | |
| Stage, | – | – | |
| I | 110 (36.4) | ||
| II | 66 (21.9) | ||
| III | 53 (17.5) | ||
| IV | 73 (24.2) | ||
| T, | – | – | |
| 1 | 113 (37.4) | ||
| 2 | 63 (20.9) | ||
| 3 | 59 (19.5) | ||
| 4 | 67 (22.2) | ||
| N, | – | – | |
| 0 | 252 (83.4) | ||
| 1 | 17 (5.6) | ||
| 2 | 33 (10.9) | ||
| M, | – | – | |
| 0 | 299 (99.0) | ||
| 1 | 2 (0.7) | ||
| 2 | 1 (0.3) | ||
| G, | – | – | |
| 0 | 1 (0.3) | ||
| 1 | 89 (29.5) | ||
| 2 | 187 (61.9) | ||
| 3 | 25 (8.3) | ||
*Pearson Chi-Square, **Student’s
LSCC group. A detailed otorhinolaryngological examination, including flexible endoscopy and/or video laryngostroboscopy, and neck palpation was performed for all LSCC patients at the Out-patient Office in the Department of Otorhinolaryngology, LUHS. Peripheral venous blood samples were collected before preparation for general anesthesia and direct microlaryngoscopy with biopsy. The diagnosis of LSCC was histologically confirmed at the Department of Pathology, LUHS. The final diagnosis of LSCC was based on clinical data and the results of histological examination and laryngeal and neck CT or MRI data. The staging of LSCC was accomplished as described by the American Joint Committee on Cancer [35].
Healthy controls. Subjects who were consulted at the Out-patient Office of the Department of Otorhinolaryngology, LUHS, and selected for surgical treatment (tympanoplasty, ossiculoplasty, tympanostomy, nasal bones reposition, septoplasty, rhinoseptoplasty, uvulopalatopharyngoplasty, or radiofrequency thermoablation of soft palate) for disorders not associated with the development of LSCC or other cancerogenic disease were enrolled in the present study as a healthy control group. The peripheral venous blood samples from these patients were collected from the same catheter that was used to induce general anesthesia. Individuals with no diagnosed oncological diseases who attended the family doctor’s consultation for general health check-ups and had a peripheral blood sample collection were also included in this study as healthy controls. All patients with the diagnosed another type and localization of cancer, acute or chronic infectious disease, individuals using psychomotor suppressants and anti-epileptic drugs, and persons younger than 18 years old were excluded from this study.
Survival rate. Data on the LSCC group mortality rate, including the survival period after diagnosis of LSCC and the cause of death, were collected from the Lithuanian State Register of Death Cases and Their Causes.
2.3DNA extraction, genotyping, and enzyme-linked immunosorbent assay
We extracted DNA samples from peripheral venous blood using the DNA salting-out method. The genotyping of all three SNPs was performed using TaqMan
2.4Quality control of genotyping
The repetitive analysis of 5% randomly chosen samples was performed for all SNPs to confirm the same rate of genotypes from initial and repetitive genotyping.
2.5Statistical analysis
Statistical analysis was performed applying the SPSS/W 20.0 software (Statistical Package for the Social Sciences for Windows, Inc., Chicago, Illinois, USA).
The data of subjects’ age were presented as the mean with standard deviation (SD) and median with interquartile range (IQR). The Student
3.Results
The distributions of the analyzed SNPs (SIRT1 rs3818292, rs3758391, and rs7895833) in the control group matched the Hardy-Weinberg equilibrium (HWE) (
Table 2
Analysis of Hardy-Weinberg equilibrium in the control group
| SNP | Allele frequencies | Genotype distribution | ||
|---|---|---|---|---|
| SIRT1 rs3818292 | 0.93 A | 0.07 G | 3/49/357 | 0.364 |
| SIRT1 rs3758391 | 0.75 C | 0.25 T | 18/169/222 | 0.043 |
| SIRT1 rs7895833 | 0.86 A | 0.14 G | 5/106/298 | 0.190 |
We analyzed the distributions of frequencies of the following genotypes and alleles of SIRT1 rs3818292, rs3758391, and rs7895833 in the LSCC and control groups. There was no statistically significant difference between these two groups (Table 3). Binomial logistic regression was performed to evaluate the impact of SIRT1 rs3818292, rs3758391, and rs7895833 on LSCC development. Our results revealed that SIRT1 rs3758391T/T genotype carriers had statistically significantly increased LSCC development odds under the codominant model (OR
Table 3
Frequencies of genotypes and alleles of SIRT1 rs3818292, rs3758391 and rs7895833 in patients with LSCC and the control group
| Polymorphism | LSCC | Control group | |
|---|---|---|---|
| rs3818292 | |||
| A/A | 255 (84.4) | 357 (87.3) | 0.133 |
| A/G | 47 (15.6) | 49 (12.0) | |
| G/G | 0 (0) | 3 (7.0) | |
| Total | 302 (100) | 409 (100) | |
| Allele | |||
| A | 557 (92.2) | 763 (93.3) | 0.445 |
| G | 47 (7.8) | 55 (6.7) | |
| rs3758391 | |||
| C/C | 151 (50.0) | 222 (54.3) | 0.115 |
| C/T | 127 (42.1) | 169 (41.3) | |
| T/T | 24 (7.9) | 18 (4.4) | |
| Total | 302 (100) | 409 (100) | |
| Allele | |||
| C | 429 (71.0) | 613 (75.3) | 0.099 |
| T | 175 (28.0) | 205 (24.7) | |
| rs7895833 | |||
| A/A | 215 (71.2) | 298 (72.9) | 0.819 |
| A/G | 82 (27.2) | 106 (25.9) | |
| G/G | 5 (1.7) | 5 (1.2) | |
| Total | 302 (100) | 409 (100) | |
| Allele | |||
| A | 512 (84.8) | 702 (85.8) | 0.579 |
| G | 92 (15.2) | 116 (14.18) |
Table 4
Binomial logistic regression analysis of SIRT1 rs3818292, rs3758391, and rs7895833 in the control and patients with LSCC groups
| Model | Genotype/allele | OR (95% CI) | AIC | |
|---|---|---|---|---|
| SIRT1 rs3818292 | ||||
| Codominant | A/G vs. A/A | 1.343 (0.872–2.067) | 0.180 | 968.375 |
| G/G vs. A/A | 0.000 (0.000–0.000) | 0.999 | ||
| Dominant | A/G | 1.265 (0.826–1.937) | 0.279 | 970.323 |
| Recessive | G/G vs. A/A | 0.000 (0.000–0.000) | 0.999 | 968.164 |
| Overdominant | A/G vs. A/A | 1.354 (0.880–2.084) | 0.168 | 969.599 |
| Additive | G | 1.173 (0.781–1.763) | 0.442 | 970.903 |
| SIRT1 rs3758391 | ||||
| Codominant | C/T vs. C/C | 1.105 (0.811–1.505) | 0.528 | 969.216 |
| T/T vs. C/C | 1.960 (1.028–3.737) | 0.041 | ||
| Dominant | C/T | 1.187 (0.881–1.599) | 0.259 | 970.216 |
| Recessive | T/T vs. C/C | 1.875 (0.999–3.522) | 0.051 | 967.617 |
| Overdominant | C/T vs. C/C | 1.031 (0.762–1.393) | 0.845 | 971.453 |
| Additive | T | 1.237 (0.968–1.581) | 0.089 | 968.600 |
| SIRT1 rs7895833 | ||||
| Codominant | A/G vs. A/A | 1.072 (0.765–1.503) | 0.685 | 973.095 |
| G/G vs. A/A | 1.386 (0.396–4.847) | 0.609 | ||
| Dominant | A/G | 1.086 (0.780–1.513) | 0.624 | 971.251 |
| Recessive | G/G vs. A/A | 1.360 (0.390–4.741) | 0.629 | 971.259 |
| Overdominant | A/G vs. A/A | 1.065 (0.761–1.492) | 0.712 | 971.355 |
| Additive | G | 1.093 (0.805–1.484) | 0.568 | 971.166 |
OR: odds ratio; CI: confidence interval;
We analyzed the distributions of SIRT1 rs3818292, rs3758391, and rs7895833 genotype and allele frequencies between the control and patient with LSCC (in early- and advanced-stages) groups. The patient group was divided into subgroups: early (I–II stage) and advanced (III–IV stage) LSCC. The SIRT1 rs3758391 TT genotype and SIRT1 rs7895833 allele G were statistically significantly more frequent in the advanced LSCC stage subgroup than in the control group (9.5% vs. 4.4%,
Table 5
Frequencies of genotypes and alleles of SIRT1 rs3818292, rs3758391, and rs7895833 in the control group and early and advanced stage LSCC subgroups
| Genotype/allele | Control group % | Early-stage (I | Advanced stage | |||
|---|---|---|---|---|---|---|
| (III | ||||||
| rs3818292 | ||||||
| A/A | 357 (87.3) | 152 (86.4) | 0.455 | 103 (81.7) | 0.130 | 0.275 |
| A/G | 49 (12.0) | 24 (13.6) | 23 (18.3) | |||
| G/G | 3 (0.7) | 0 (0.0) | 0 (0.0) | |||
| Total | 409 (100) | 176 (100) | 126 (100) | |||
| Allele | ||||||
| A | 763 (93.3) | 328 (93.2) | 0.953 | 229 (90.9) | 0.199 | 0.296 |
| G | 55 (6.7) | 24 (8.6) | 23 (9.1) | |||
| rs3758391 | ||||||
| C/C | 222 (54.3) | 89 (50.6) | 0.413 | 62 (49.2) | 0.083 | 0.692 |
| C/T | 169 (41.3) | 75 (42.6) | 52 (41.3) | |||
| T/T | 18 (4.4) | 12 (6.8) | 12 (9.5) | |||
| Total | 409 (100) | 176 (100) | 126 (100) | |||
| Allele | ||||||
| C | 613 (74.9) | 253 (71.9) | 0.273 | 176 (69.8) | 0.108 | 0.587 |
| T | 205 (25.1) | 99 (28.1) | 76 (30.2) | |||
| rs7895833 | ||||||
| A/A | 298 (72.9) | 132 (75.0) | 0.706 | 83 (65.9) | 0.154 | 0.081 |
| A/G | 106 (25.9) | 43 (24.4) | 39 (31.0) | |||
| G/G | 5 (1.2) | 1 (0.6) | 4 (3.2) | |||
| Total | 409 (100) | 176 (100) | 126 (100) | |||
| Allele | ||||||
| A | 702 (85.8) | 307 (87.2) | 0.525 | 205 (81.3) | 0.084 | 0.048 |
| G | 116 (14.2) | 45 (12.8) | 47 (18.7) |
*Early stage vs. advanced stage;
Table 6
Binomial logistic regression analysis of SIRT1 rs3758391 in healthy controls and patients with LSCC advanced stage
| Model | Genotype/allele | OR (95% CI) | AIC | |
|---|---|---|---|---|
| SIRT1 rs3758391 | ||||
| Codominant | C/T vs. C/C | 1.102 (0.724–1.676) | 0.651 | 583.597 |
| T/T vs. C/C | 2.387 (1.091–5.222) | 0.029 | ||
| Dominant | C/T | 1.225 (0.822–1.828) | 0.319 | 585.070 |
| Recessive | T/T vs. C/C | 2.287 (1.070–4.888) | 0.033 | 581.802 |
| Overdominant | C/T vs. C/C | 0.998 (0.665–1.497) | 0.992 | 586.063 |
| Additive | T | 1.317 (0.951–1.823) | 0.097 | 583.341 |
OR: odds ratio; CI: confidence interval;
The LSCC patients’ group was divided into four subgroups according to tumor size (T1, T2, T3, and T4). Analysis of SIRT1 rs3818292, rs3758391, and rs7895833 genotypes and allele frequencies between the control group and patients’ subgroups was performed. SIRT1 rs3758391 TT genotype was more frequent in T4 subgroup than in controls (11.9% vs. 4.4%,
Table 7
Frequencies of genotypes and alleles of SIRT1 rs3818292, rs3758391, and rs7895833 in the healthy control group and LSCC patients with different tumor size
| Genotype/allele | Control group % | T1 % | T2 % | T3 % | T4 % | ||||
|---|---|---|---|---|---|---|---|---|---|
| rs3818292 | |||||||||
| A/A | 357 (87.3) | 95 (84.1) | 0.366 | 57 (90.5) | 0.668 | 46 (78.0) | 0.087 | 57 (85.1) | 0.629 |
| A/G | 49 (12.0) | 18 (15.9) | 6 (9.5) | 13 (22.0) | 10 (14.9) | ||||
| G/G | 3 (0.7) | 0 (0.0) | 0 (0.0) | 0 (0.0) | 0 (0.0) | ||||
| Total | 409 (100) | 113 (100) | 63 (100) | 59 (100) | 67 (100) | ||||
| Allele | |||||||||
| A | 763 (93.3) | 208 (92.0) | 0.517 | 120 (95.2) | 0.404 | 105 (89.0) | 0.093 | 124 (92.5) | 0.753 |
| G | 55 (6.7) | 18 (8.0) | 6 (4.8) | 13 (11.0) | 10 (7.5) | ||||
| rs3758391 | |||||||||
| C/C | 222 (54.3) | 54 (47.8) | 0.217 | 35 (55.6) | 0.967 | 30 (50.8) | 0.688 | 32 (47.8) | 0.039 |
| C/T | 169 (41.3) | 50 (44.2) | 25 (39.7) | 25 (42.4) | 27 (40.3) | ||||
| T/T | 18 (4.4) | 9 (8.0) | 3 (4.8) | 4 (6.8) | 8 (11.9) | ||||
| Total | 409 (100) | 113 (100) | 63 (100) | 59 (100) | 67 (100) | ||||
| Allele | |||||||||
| C | 613 (74.9) | 158 (69.9) | 0.128 | 95 (75.4) | 0.912 | 85 (72.0) | 0.459 | 91 (67.9) | 0.086 |
| T | 205 (25.1) | 68 (30.1) | 31 (24.6) | 33 (28.0) | 43 (32.1) | ||||
| rs7895833 | |||||||||
| A/A | 298 (72.9) | 84 (74.3) | 0.494 | 49 (77.8) | 0.657 | 36 (61.0) | 0.085 | 46 (68.7) | 0.030 |
| A/G | 106 (25.9) | 29 (25.7) | 13 (20.6) | 23 (39.0) | 17 (25.4) | ||||
| G/G | 5 (1.2) | 0 (0.0) | 1 (1.6) | 0 (0.0) | 4 (6.0) | ||||
| Total | 409 (100) | 113 (100) | 188 (100) | 59 (100) | 67 (100) | ||||
| Allele | |||||||||
| A | 702 (85.8) | 197 (87.2) | 0.604 | 111 (88.1) | 0.491 | 95 (80.5) | 0.129 | 109 (81.3) | 0.176 |
| G | 116 (14.2) | 29 (12.8) | 15 (11.9) | 23 (19.5) | 25 (18.7) |
T – tumor size;
Table 8
Binomial logistic regression analysis of SIRT1 rs3818292, rs7895833 in healthy control and patients with LSCC T3 groups
| Model | Genotype/allele | OR (95% CI) |
| AIC |
|---|---|---|---|---|
| SIRT1 rs3818292 | ||||
| Codominant | A/G vs. A/A | 2.059 (1.039–4.081) | 0.039 | 353.882 |
| G/G vs. A/A | 0.000 (0.000–0.000) | 0.999 | ||
| Dominant | A/G | 1.940 (0.982–3.833) | 0.056 | 353.257 |
| Recessive | G/G vs. A/A | 0.000 (0.000–0.000) | 0.999 | 355.787 |
| Overdominant | A/G vs. A/A | 2.076 (1.048–4.115) | 0.036 | 352.606 |
| Additive | G | 1.703 (0.903–3.214) | 0.100 | 354.123 |
| SIRT1 rs7895833 | ||||
| Codominant | A/G vs. A/A | 1.796 (1.018–3.170) | 0.043 | 353.311 |
| G/G vs. A/A | 0.000 (0.000–0.000) | 0.999 | ||
| Dominant | A/G | 1.715 (0.973–3.023) | 0.062 | 353.234 |
| Recessive | G/G vs. A/A | 0.000 (0.000–0.000) | 0.999 | 355.243 |
| Overdominant | A/G vs. A/A | 1.826 (1.035–3.223) | 0.038 | 352.442 |
| Additive | G | 1.531 (0.902–2.597) | 0.114 | 354.208 |
OR: odds ratio; CI: confidence interval;
Table 9
Binomial logistic regression analysis of SIRT1 rs3758391, rs7895833 in healthy control and patients with LSCC T4 groups
| Model | Genotype/allele | OR (95% CI) |
| AIC |
|---|---|---|---|---|
| SIRT1 rs3758391 | ||||
| Codominant | C/T vs. C/C | 1.108 (0.640–1.921) | 0.714 | 385.607 |
| T/T vs. C/C | 3.083 (1.239–7.672) | 0.015 | ||
| Dominant | C/T | 1.298 (0.774–2.178) | 0.322 | 357.850 |
| Recessive | T/T vs. C/C | 2.945 (1.226–7.077) | 0.016 | 383.741 |
| Overdominant | C/T vs. C/C | 0.959 (0.5661.623) | 0.875 | 388.805 |
| Additive | T | 1.454 (0.961–2.201) | 0.077 | 385.760 |
| SIRT1 rs7895833 | ||||
| Codominant | A/G vs. A/A | 1.039 (0.571–1.891) | 0.90 | 385.843 |
| G/G vs. A/A | 5.183 (1.342–20.010) | 0.017 | ||
| Dominant | A/G | 1.226 (0.700–2.146) | 0.477 | 388.333 |
| Recessive | G/G vs. A/A | 5.130 (1.342–19.618) | 0.017 | 383.859 |
| Overdominant | A/G vs. A/A | 0.972 (0.537–1.759) | 0.925 | 388.821 |
| Additive | G | 1.401 (0.862–2.277) | 0.173 | 387.051 |
OR: odds ratio; CI: confidence interval;
We also performed a comparative analysis of the distribution of the SIRT1 rs3818292, rs3758391, and rs7895833 genotypes and alleles in the LSCC patient subgroups with metastasis to the neck lymph nodes and without metastases to the neck lymph nodes, and the control group. We found that the rs3758391 TT genotype was statistically significantly more frequent in patients without metastases to the neck lymph nodes than in healthy controls (8.3% vs. 4.4%,
Table 10
Frequencies of genotypes and alleles of SIRT1 rs3818292, rs3758391, and rs7895833 in healthy controls and LSCC patients with and without metastases to the neck lymph nodes
| Genotype/allele | Control group % | Not -spread to lymph nodes % | Spread to lymph nodes % | |||
|---|---|---|---|---|---|---|
| rs3818292 | ||||||
| A/A | 357 (87.3) | 210 (83.3) | 0.099 | 45 (90) | 0.759 | 0.235 |
| A/G | 49 (12.0) | 42 (16.7) | 5 (10.0) | |||
| G/G | 3 (0.7) | 0 (0.0) | 0 (0.0) | |||
| Total | 409 (100) | 252 (100) | 50 (100) | |||
| Allele | ||||||
| A | 763 (93.3) | 462 (91.7) | 0.276 | 95 (95.0) | 0.510 | 0.256 |
| G | 55 (6.7) | 42 (8.3) | 5 (5.0) | |||
| rs3758391 | ||||||
| C/C | 222 (54.3) | 124 (49.2) | 0.086 | 27 (54.0) | 0.874 | 0.764 |
| C/T | 169 (41.3) | 107 (42.5) | 20 (40.0) | |||
| T/T | 18 (4.4) | 21 (8.3) | 3 (6.0) | |||
| Total | 409 (100) | 252 (100) | 50 (100) | |||
| Allele | ||||||
| C | 613 (74.9) | 355 (70.4) | 0.073 | 74 (74.0) | 0.838 | 0.473 |
| T | 205 (25.1) | 149 (29.6) | 26 (26.0) | |||
| rs7895833 | ||||||
| A/A | 298 (72.9) | 178 (70.6) | 0.817 | 37 (74.0) | 0.282 | 0.267 |
| A/G | 106 (25.9) | 71 (28.2) | 11 (22.0) | |||
| G/G | 5 (1.2) | 3 (1.2) | 2 (4.0) | |||
| Total | 409 (100) | 252 (100) | 57 (100) | |||
| Allele | ||||||
| A | 702 (85.8) | 427 (84.7) | 0.583 | 85 (85.0) | 0.825 | 0.944 |
| G | 116 (14.2) | 77 (15.3) | 15 (15.0) |
*Metastases to the neck lymph nodes vs. no metastases;
The LSCC patient group was divided into three subgroups according to the tumor differentiation grade G1, G2, and G3. Analysis of SIRT1 rs3818292, rs3758391, and rs7895833 genotypes and allele frequencies between the control group and patient subgroups was performed. The SIRT1 rs3758391 TT genotype, as well as T allele, were more frequent in the G1 subgroup than in controls (10.0% vs. 4.4%,
Table 11
Frequencies of genotypes and alleles of SIRT1 rs3818292, rs3758391, and rs7895833 in healthy controls and LSCC patients with different tumor differentiation grades
| Genotype/allele | Control group % | G1 % | G2 % | G3 % | |||
|---|---|---|---|---|---|---|---|
| rs3818292 | |||||||
| A/A | 357 (87.3) | 73 (81.1) | 0.161 | 162 (86.2) | 0.417 | 21 (84.0) | 0.769 |
| A/G | 49 (12.0) | 17 (18.9) | 26 (13.8) | 4 (16.0) | |||
| G/G | 3 (0.7) | 0 (0.0) | 0 (0.0) | 0 (0.0) | |||
| Total | 409 (100) | 90 (100) | 188 (100) | 25 (100) | |||
| Allele | |||||||
| A | 763 (93.3) | 163 (90.6) | 0.201 | 350 (93.1) | 0.903 | 46 (92.0) | 0.728 |
| G | 55 (6.7) | 17 (9.4) | 26 (6.9) | 4 (8.0) | |||
| rs3758391 | |||||||
| C/C | 222 (54.3) | 41 (45.6) | 0.064 | 99 (52.7) | 0.857 | 12 (48.0) | 0.003 |
| C/T | 169 (41.3) | 40 (44.4) | 79 (42.0) | 8 (32.0) | |||
| T/T | 18 (4.4) | 9 (10.0) | 10 (5.3) | 5 (20.0) | |||
| Total | 409 (100) | 90 (100) | 188 (100) | 25 (100) | |||
| Allele | |||||||
| C | 613 (74.9) | 122 (67.8) | 0.048 | 277 (73.7) | 0.64 | 32 (64.0) | 0.086 |
| T | 205 (25.1) | 58 (32.2) | 99 (26.3) | 18 (36.0) | |||
| rs7895833 | |||||||
| A/A | 298 (72.9) | 61 (67.8) | 0.602 | 140 (74.5) | 0.705 | 15 (60.0) | |
| A/G | 106 (25.9) | 28 (31.1) | 47 (25.0) | 7 (28.0) | |||
| G/G | 5 (1.2) | 1 (1.1) | 1 (0.5) | 3 (12.0) | |||
| Total | 409 (100) | 90 (100) | 188 (100) | 3 (100) | |||
| Allele | |||||||
| A | 702 (85.8) | 150 (83.3) | 0.393 | 327 (87.0) | 0.493 | 37 (74.0) | 0.023 |
| G | 116 (14.2) | 30 (16.7) | 49 (13.0) | 13 (26.0) |
G-Tumor differentiation grade;
SIRT1 serum levels were measured in 24 samples from the control group and 29 from the LSCC group. We did not find any statistically significant differences in SIRT1 serum levels between the LSCC and control groups (0.395 (0.714) ng/ml vs. 0.417 (0.596) ng/ml,
Figure 1.
Serum SIRT1 levels in controls and patients with LSCC.
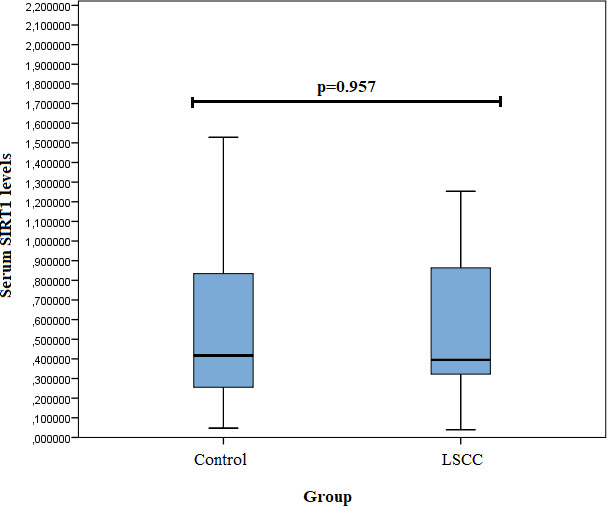
Table 12
Genotype distribution and serum SIRT1 levels
| SNP/genotype | SIRT1 level (pg/mL) | ||
| LSCC median (IQR) | Controls median (IQR) | ||
| SIRT1 rs3818292 | |||
| A/A | 0.48 (0.86) | 0.42 (0.62) | 0.396* |
| A/G | 0.20 (0.30) | 0.57 ( | 0.034* |
| G/G | – | – | – |
| SIRT1 rs3758391 | |||
| C/C | 0.39 (0.57) | 0.36 (0.42) | 0.661* |
| C/T | 0.56 (1.33) | 0.65 (2.01) | 0.622* |
| T/T | – | – | – |
| SIRT1 rs7895833 | |||
| A/A | 0.40 (0.76) | 0.41 (0.52) | 0.587* |
| A/G | 0.39 (4.92) | 0.65 (14.18) | 0.338* |
| G/G | – | – | – |
* Obtained using the Mann-Whitney
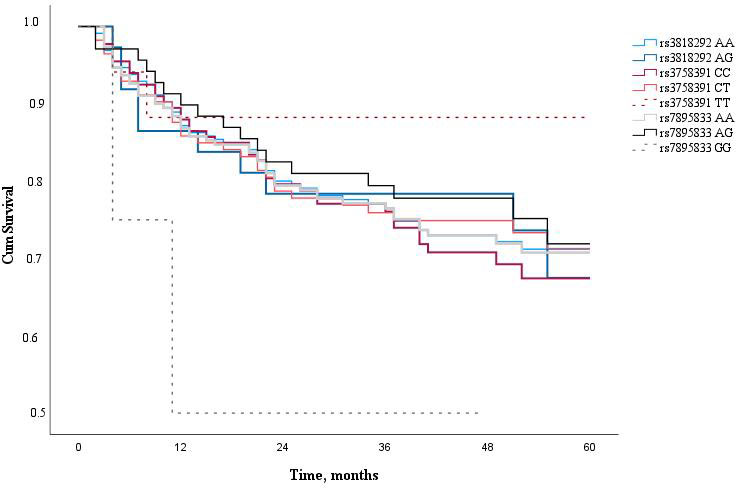
Finally, the 5-year survival rate of LSCC patients was determined. The 5-year overall survival rate of LSCC patients was 62.7% and the LSCC-specific survival rate of five years was 72.6%. Analyzing LSCC patients’ 5-year survival rate and the genotype distribution of SIRT1 (rs3818292, rs3758391, and rs7895833), we have not found a statistically significant impact of these SNPs on the patients’ 5-year survival rate (Fig. 2).
Figure 2.
Five year survival analysis of LSCC patients according to SIRT1 SNPs (rs3818292, rs3758391, and rs7895833).
4.Discussion
The SIRT1 is located on the long arm of chromosome 10 (10q21.3) [36]. Evidence from many studies has revealed that SIRT1 participates in the regulation of diverse cellular processes, including carcinogenesis [37]. In cancer development, the role of SIRT1 is dual because it can function as both a tumor promoter and a tumor suppressor [28, 37, 38, 39]. The SIRT1 acts as a tumor suppressor in holding genomic stability and negatively regulates anti-apoptotic genes, resulting in increased anticancer ability [37]. On the other hand, to meet the tumor’s high-energy needs and promote its development, SIRT1 not only induces angiogenesis but also improves HIF1
Epigenetic modifications are critical for LSCC carcinogenesis [8]. Gene variants may account for different SIRT1 expressions, determining individuals’ susceptibility to specific pathologies [19, 24, 26, 40, 41, 42, 43, 44, 45]. However, little is known about the effect of polymorphism of the SIRT1 in both LSCC carcinogenesis and disease progression. Therefore, the focus of the present study was to estimate the associations between SIRT1 SNPs (rs3818292, rs3758391, rs7895833), SIRT1 serum levels, and LSCC.
In the present study, we found that SIRT1 rs3758391 T/T genotype carriers had statistically significantly increased odds of LSCC. Moreover, the current study results revealed that patients with this genotype had 2.387 odds of LSCC development into advanced stages. To the best of our knowledge, this is the first report that associates SNP in the SIRT1 gene (rs3758391) with LSCC in a pure and homogenous LSCC patients’ cohort.
According to the literature, the SRT1 rs3758391 polymorphism has been associated with healthy aging in the Chinese population and better cognitive function in older individuals [46, 47, 48]. Also, several studies have shown the role of this SIRT1 SNP in several diseases, including type 2 diabetes mellitus [49], cardiovascular diseases [50], systemic lupus erythematosus [51], and major depressive disorder [52]. Also, the accumulating evidence has suggested the importance of the SIRT1 rs3758391 in cancer development [43, 45, 53, 54]. Furthermore, Shafieian et al. found that the SIRT1 rs3758391 TT genotype increased the risk of urinary bladder cancer in the Iranian population [45]. Shaker et al. study showed that carriers of the SIRT1 rs3758391 CC genotype were more often in the colorectal cancer group than in the control group [54]. Rizk et al. study revealed that the SIRT rs3758391 polymorphism (TT genotype) was associated with the increased breast cancer risk and that the SIRT1 rs3758391T allele was a potential risk factor for breast cancer [43]. Kan et al. conducted a large-scale case-control study on diffuse large B cell lymphoma and compared results of patients with the healthy control group. They found that the SIRT1 rs3758391T allele presented an increased risk of diffuse large B cell lymphoma. Also, they noticed SIRT1 mRNA expression upregulation in diffuse large B cell lymphoma patients with TT genotype. Moreover, a worse survival rate was revealed in patients with TT genotypes. The authors concluded that the SIRT1 rs3758391 might be used as a biomarker for the prognosis of susceptibility and survival of diffuse large B cell lymphoma [53]. We did not find a significant impact of SIRT1 SNPs on LSCC patients’ 5-year survival rate in the present study. Lv et al. did not find any association between SIRT1 gene polymorphisms, including rs3758391, and lung cancer [55].
The results of the present study revealed that two SNPs of SIRT1 (rs3818292 and rs7895833) are associated with the increased odds of LSCC development into T3 and T4. No associations between the aforementioned SIRT1 SNPs and any cancer were described in the literature, so far. However, these polymorphisms of SIRT1 have been investigated mainly in the context of metabolism. They have been associated with the increased risk of visceral (rs3818292) and general (rs7895833) obesity in adults and decreased risk (a protective effect) of obesity in children (rs7895833) [56, 57, 58]. Associations of SIRT1 rs3818292 with diabetic nephropathy and nondiabetic type 1 cardiorenal syndrome have also been found [59, 60]. Kilic et al., in their study, found that SIRT1 rs7895833 might affect longevity. They observed a significant increase in serum SIRT1 level in older people and carriers of SIRT1 rs7895833, suggesting an association between SIRT1 rs7895833 SNP and longevity [61].
In the present study, we measured the serum concentration of SIRT1 in patients with LSCC and compared it with the results of the control group. We did not find any statistically significant differences between these groups. However, comparing the genotype distribution and serum levels of SIRT1 between the control and LSCC groups, we found that only LSCC patients with SIRT1 rs3818292 AG genotype demonstrated a clear tendency to significantly lower SIRT1 serum levels than controls. These findings are in some contradiction with the data of the literature. For instance, Shaker et al. in their study found that patients with colorectal cancer had a significant increment in the mean level of serum SIRT1 as compared to the control group and the mean level of serum SIRT1 was higher in patients with tumor size
On the other hand, Noguchi et al. assessed the SIRT1 expression immunohistochemically using surgical samples from 437 consecutive head and neck squamous cell carcinoma (HNSCC) patients. Results of the study showed that the SIRT1 expression acted as a tumor suppressor and served as an indicator for a good prognosis [33]. Yu et al. investigated SIRT1 expression in laryngeal and hypopharyngeal squamous cell carcinoma (HSCC) tissue samples using immunohistochemical staining and its correlations with clinical parameters. They found a significant correlation between the decreased SIRT1 expression and clinical tumor stage, metastasis to the neck lymph nodes, and the patient’s survival rate. The authors concluded that SIRT1 acts as a tumor suppressor in LSCCs and HSCC [64]. However, those authors pulled their LSCC and HNSCC patients into one group; therefore, these results cannot be directly compared to the cohort of pure LSCC patients used in the present study. On the other hand, the results of the current study may suggest that in cases of LSCC, the SIRT1 serum levels remain unchanged despite the increment of SIRT1 expression directly in the tumor tissue. This phenomenon may be caused by a relatively low tumor suppressor activity of SIRT1 related to the less aggressive nature of LSCC.
The strength of the present study is a relatively large and homogenous group of pure LSCC patients (302 subjects) and age and sex-matched controls (409 subjects). This specific feature ensured a comprehensive analysis of associations between SIRT1 serums levels and selected SIRT1 SNPs and the development of a particular tumor in one anatomical region of the head and neck, i.e., LSCC.
Several limitations of the present study must be considered. The sample size for analysis of serum concentration levels of SIRT1 was limited and too small to reach the desired power setting. Therefore, the results obtained in the present study regarding the associations between the SIRT1 serum concentration levels and LSCC should be considered as the tendency. Further investigation with a large enough sample size is foreseen to confirm the possible role of serum concentration levels of SIRT1 in LSCC development. The data about smoking and alcohol consumption was available and analyzed in a limited cohort of study participants that did not match the patient and control groups. Therefore, the impact of smoking and alcohol consumption habits was not completely investigated. However, this is the target task for future investigation.
5.Conclusion
The present study showed a statistically significant association between SIRT1 rs3758391T/T genotype and increased odds of LSCC development, as well as increased LSCC development into the advanced stages and T4 odds. Also, SIRT1 rs3818292 and rs7895833 SNPs revealed increased odds of LSCC development into T3 and T4. LSCC patients with SIRT1 rs3818292 AG genotype showed a tendency to manifest with lower SIRT1 serum levels. No associations between SIRT1 (rs3818292, rs3758391, and rs7895833) SNPs and the 5-year survival rate of LSCC patients were found in the present study. Therefore, the role of SIRT1 in cancer, including LSCC, remains incompletely understood. Further research is needed to investigate possible mechanisms of the associations found, before selecting candidates for prognostic factors or a future therapeutic target.
Author contributions
Conception: R.L., V.U.
Interpretation or analysis of data: R.L., A.V., G.G., V.U., P.V., V.L.
Preparation of the manuscript: P.V., V.U.
Revision for important intellectual content: R.L., V.U., V.L.
Supervision: R.L., V.U.
References
[1] | World Health Organization, Cancer Tomorrow, (2020) , 1–2. https://gco.iarc.fr/today/data/factsheets/cancers/14-Larynx-fact-sheet.pdf. |
[2] | World Health Organization, and International agency for research on cancer, Estimated number of deaths from 2020 to 2040, Both sexes, age [0–85+], (2020) . https://gco.iarc.fr/tomorrow/en/dataviz/bubbles?sexes=0&mode=population&group_populations=0&multiple_cancers=1&cancers=39_14&group_cancers=1. |
[3] | G. Di Credico, J. Polesel, L. Dal Maso, F. Pauli, N. Torelli, D. Luce, L. Rado, K. Matsuo, D. Serraino, P. Brennan, I. Holcatova, W. Ahrens, P. Lagiou, C. Canova, L. Richiardi, C.M. Healy, K. Kjaerheim, D.I. Conway, G.J. Macfarlane, P. Thomson, A. Agudo, A. Znaor, S. Franceschi, R. Herrero, T.N. Toporcov, R.A. Moyses, J. Muscat, E. Negri, M. Vilensky, L. Fernandez, M.P. Curado, A. Menezes, A.W. Daudt, R. Koifman, V. Wunsch-Filho, A.F. Olshan, J.P. Zevallos, E.M. Sturgis, G. Li, F. Levi, Z.F. Zhang, H. Morgenstern, E. Smith, P. Lazarus, C. La Vecchia, W. Garavello, C. Chen, S.M. Schwartz, T. Zheng, T.L. Vaughan, K. Kelsey, M. McClean, S. Benhamou, R.B. Hayes, M.P. Purdue, M. Gillison, S. Schantz, G.P. Yu, S.C. Chuang, P. Boffetta, M. Hashibe, A.L. Yuan-Chin and V. Edefonti, Alcohol drinking and head and neck cancer risk: The joint effect of intensity and duration, Br J Cancer 123: ((2020) ), 1456–1463. doi: 10.1038/s41416-020-01031-z. |
[4] | K.R. Sterba, E. Garrett-mayer, M.J. Carpenter, J.A. Tooze, J.L. Hatcher, C. Sullivan, L. Anne, G.W. Warren, T.A. Day, A.J. Alberg and K.E. Weaver, Smoking status and symptom burden in surgical head and neck cancer patients, Laryngoscope 127: ((2018) ), 127–133. doi: 10.1002/lary.26159.Smoking. |
[5] | N.K. Proia, G.M. Paszkiewicz, M.A.S. Nasca, G.E. Franke and J.L. Pauly, Smoking and smokeless tobacco-associated human buccal cell mutations and their association with oral cancer – A review, Cancer Epidemiol Biomarkers Prev 15: ((2006) ), 1061–1077. doi: 10.1158/1055-9965.EPI-05-0983. |
[6] | O. Bayer, R. Cmara, S.R. Zeissig, M. Ressing, A. Dietz, L.D. Locati, H. Ramroth and S. Singer, Occupation and cancer of the larynx: A systematic review and meta-analysis, Eur Arch Oto-Rhino-Laryngology 273: ((2016) ), 9–20. doi: 10.1007/s00405-014-3321-y. |
[7] | T. Ramsey, E. Guo, P.F. Svider, H. Lin, S. Syeda, S.N. Raza and A.M. Fribley, Laryngeal cancer: Global socioeconomic trends in disease burden and smoking habits, Laryngoscope 128: ((2018) ), 2039–2053. doi: 10.1002/lary.27068. |
[8] | J. Gaździcka, K. Gołąbek, J.K. Strzelczyk and Z. Ostrowska, Epigenetic modifications in head and neck cancer, Biochem Genet 58: ((2020) ), 213–244. doi: 10.1007/s10528-019-09941-1. |
[9] | A. Dietz, H. Ramroth, T. Urban, W. Ahrens and H. Becher, Exposure to cement dust, related occupational groups and laryngeal cancer risk: Results of a population based case-control study, Int J Cancer 108: ((2004) ), 907–911. doi: 10.1002/ijc.11658. |
[10] | N. Shivappa, J.R. Hbert, V. Rosato, D. Serraino and C. La Vecchia, Inflammatory potential of diet and risk of laryngeal cancer in a case–control study from Italy, Cancer Causes Control 27: ((2016) ), 1027–1034. doi: 10.1007/s10552-016-0781-y. |
[11] | H. Becher, H. Ramroth, W. Ahrens, A. Risch, P. Schmezer and A. Dietz, Occupation, exposure to polycyclic aromatic hydrocarbons and laryngeal cancer risk, Int J Cancer 116: ((2005) ), 451–457. doi: 10.1002/ijc.21049. |
[12] | H. Ramroth, W. Ahrens, A. Dietz and H. Becher, Occupational asbestos exposure as a risk factor for laryngeal carcinoma in a population-based case-control study from Germany, Am J Ind Med 54: ((2011) ), 510–514. doi: 10.1002/ajim.20963. |
[13] | O. Shangina, P. Brennan, N. Szeszenia-Dabrowska, D. Mates, E. Fabinov, T. Fletcher, A. T’Mannetje, P. Boffetta and D. Zaridze, Occupational exposure and laryngeal and hypopharyngeal cancer risk in Central and Eastern Europe, Am J Epidemiol 164: ((2006) ), 367–375. doi: 10.1093/aje/kwj208. |
[14] | H. Ramroth, A. Dietz, W. Ahrens and H. Becher, Occupational wood dust exposure and the risk of laryngeal cancer: A population based case-control study in Germany, Am J Ind Med 51: ((2008) ), 648–655. doi: 10.1002/ajim.20605. |
[15] | C.I. Cornean, M. Cosgarea, A. Cătană, C.A. Mogoantă, V. Necula and A.A. Maniu, Do we know enough about the genetic involvement in laryngeal cancer susceptibility and prognostic outcome? Rom J Morphol Embryol 60: ((2019) ), 353–367. |
[16] | C.E. Steuer, M. El-Deiry, J.R. Parks, K.A. Higgins and N.F. Saba, An update on larynx cancer, CA Cancer J Clin 67: ((2017) ), 31–50. doi: 10.3322/caac.21386. |
[17] | R. Nocini, G. Molteni, C. Mattiuzzi and G. Lippi, Updates on larynx cancer epidemiology, Chinese J Cancer Res 32: ((2020) ), 18–25. doi: 10.21147/j.issn.1000-9604.2020.01.03. |
[18] | Y. Zhang, K. Thai, T. Jin, M. Woo and R.E. Gilbert, SIRT1 activation attenuates α cell hyperplasia, hyperglucagonaemia and hyperglycaemia in STZ-diabetic mice, Sci Rep 8: ((2018) ), 1–10. doi: 10.1038/s41598-018-32351-z. |
[19] | C. Sardu, G. Pieretti, N. D’Onofrio, F. Ciccarelli, P. Paolisso, M.B. Passavanti, R. Marfella, M. Cioffi, P. Mone, A.M. Dalise, F. Ferraraccio, I. Panarese, A. Gambardella, N. Passariello, M.R. Rizzo, M.L. Balestrieri, G. Nicoletti and M. Barbieri, Inflammatory cytokines and SIRT1 levels in subcutaneous abdominal fat: Relationship with cardiac performance in overweight pre-diabetics patients, Front Physiol 9: ((2018) ), 1–18. doi: 10.3389/fphys.2018.01030. |
[20] | M. Wojcik, K. Mac-Marcjanek and L. Wozniak, Physiological and pathophysiological functions of SIRT1, Mini-Reviews Med Chem 9: ((2012) ), 386–394. doi: 10.2174/1389557510909030386. |
[21] | C. Brochier, G. Dennis, M.A. Rivieccio, K. McLaughlin, G. Coppola, R.R. Ratan and B. Langley, Specific acetylation of p53 by HDAC inhibition prevents DNA damage-induced apoptosis in neurons, J Neurosci 33: ((2013) ), 8621–8632. doi: 10.1523/JNEUROSCI.5214-12.2013. |
[22] | C. Xu, L. Wang, P. Fozouni, G. Evjen, V. Chandra, J. Jiang, C. Lu, M. Nicastri, C. Bretz, J.D. Winkler, R. Amaravadi, B.A. Garcia, P.D. Adams, M. Ott, W. Tong, T. Johansen, Z. Dou and S.L. Berger, SIRT1 is downregulated by autophagy in senescence and ageing, Nat Cell Biol 22: ((2020) ), 1170–1179. doi: 10.1038/s41556-020-00579-5. |
[23] | T.M. Robinette, J.W. Nicholatos, A.B. Francisco, K.E. Brooks, R.Y. Diao, S. Sorbi, V. Ricca, B. Nacmias, M.A. Brieo-Enrquez and S. Libert, SIRT1 accelerates the progression of activity-based anorexia, Nat Commun 11: ((2020) ), 1–10. doi: 10.1038/s41467-020-16348-9. |
[24] | D. Nan, C. Yuqi, S. Zonglin, D. Chenglong, L. Na, L. Fang, Z. Cong and X. Xiufeng, Association of a SIRT1 polymorphism with changes of gray matter volume in patients with first-episode medication-nave major depression, Psychiatry Res – Neuroimaging 301: ((2020) ), 1–7. doi: 10.1016/j.pscychresns.2020.111101. |
[25] | Y. Zhang, S. Anoopkumar-Dukie, D. Arora and A.K. Davey, Review of the anti-inflammatory effect of SIRT1 and SIRT2 modulators on neurodegenerative diseases, Eur J Pharmacol 867: ((2020) ). doi: 10.1016/j.ejphar.2019.172847. |
[26] | S. Razi, V.C. Cogger, M. Kennerson, V.L. Benson, A.C. McMahon, F.M. Blyth, D.J. Handelsman, M.J. Seibel, V. Hirani, V. Naganathan, L. Waite, R. De Cabo, R.G. Cumming and D.G. Le Couteur, SIRT1 polymorphisms and serum-induced sirt1 protein expression in aging and frailty: The CHAMP study, Journals Gerontol – Ser A Biol Sci Med Sci 72: ((2017) ), 870–876. doi: 10.1093/gerona/glx018. |
[27] | N. D’Onofrio, L. Servillo and M.L. Balestrieri, SIRT1 and SIRT6 signaling pathways in cardiovascular disease protection, Antioxidants Redox Signal 28: ((2018) ), 711–732. doi: 10.1089/ars.2017.7178. |
[28] | H. Karbasforooshan, A. Roohbakhsh and G. Karimi, SIRT1 and microRNAs: The role in breast, lung and prostate cancers, Exp Cell Res 367: ((2018) ), 1–6. doi: 10.1016/j.yexcr.2018.03.023. |
[29] | C. Wang, W. Yang, F. Dong, Y. Guo, J. Tan, S. Ruan and T. Huang, The prognostic role of Sirt1 expression in solid malignancies: A meta-analysis, Oncotarget 8: ((2017) ), 66343–66351. doi: 10.18632/oncotarget.18494. |
[30] | M. Sun, M. Du, W. Zhang, S. Xiong, X. Gong, P. Lei, J. Zha, H. Zhu, H. Li, D. Huang and X. Gu, Survival and clinicopathological significance of SIRT1 expression in cancers: A meta-analysis, Front Endocrinol (Lausanne) 10: ((2019) ). doi: 10.3389/fendo.2019.00121. |
[31] | Z. Ye, B. Fang, J. Pan, N. Zhang, J. Huang, C. Xie, T. Lou and Z. Cao, MiR-138 suppresses the proliferation, metastasis and autophagy of non-small cell lung cancer by targeting Sirt1, Oncol Rep 37: ((2017) ), 3244–3252. doi: 10.3892/or.2017.5619. |
[32] | Y.Y. Kang, F.L. Sun, Y. Zhang and Z. Wang, SIRT1 acts as a potential tumor suppressor in oral squamous cell carcinoma, J Chinese Med Assoc 81: ((2018) ), 416–422. doi: 10.1016/j.jcma.2017.09.004. |
[33] | A. Noguchi, X. Li, A. Kubota, K. Kikuchi, Y. Kameda, H. Zheng, Y. Miyagi, I. Aoki and Y. Takano, SIRT1 expression is associated with good prognosis for head and neck squamous cell carcinoma patients, Oral Surg Oral Med Oral Pathol Oral Radiol 115: ((2013) ), 385–392. doi: 10.1016/j.oooo.2012.12.013. |
[34] | V. Uloza, T. Tamauskaite, A. Vilkeviciute, A. Pasvenskaite, V. Liutkevicius and R. Liutkeviciene, Determination of SIRT1 rs12778366, FGFR2 rs2981582, STAT3 rs744166, and RAGE rs1800625 Single Gene Polymorphisms in Patients with Laryngeal Squamous Cell Carcinoma, Dis Markers 2019: ((2019) ). doi: 10.1155/2019/3907232. |
[35] | American Joint Committee on Cancer, AJCC Cancer Staging Manual, Eighth Edition, 2018. [email protected]. |
[36] | NCBI, Gene_Result, Ncbi, (2018) . https://www.ncbi.nlm.nih.gov/gene/7054#top. |
[37] | H. Yang, Y.J. Bi, L.X. Xue, J. Wang, Y. Lu, Z.G. Zhang, X. Chen, Y. Chu, R. Yang, R. Wang and G.W. Liu, Multifaceted modulation of SIRT1 in cancer and inflammation, Crit Rev Oncog 20: ((2015) ), 49–64. doi: 10.1615/CritRevOncog.2014012374. |
[38] | S. Portmann, R. Fahrner, A. Lechleiter, A. Keogh, S. Overney, A. Laemmle, K. Mikami, M. Montani, M.P. Tschan, D. Candinas and D. Stroka, Antitumor effect of SIRT1 inhibition in human HCC tumor models in vitro and in vivo, Mol Cancer Ther 12: ((2013) ), 499–508. doi: 10.1158/1535-7163.MCT-12-0700. |
[39] | Y. Yu, Y. Liu, C. Zong, Q. Yu, X. Yang, L. Liang, F. Ye, L. Nong, Y. Jia, Y. Lu and Z. Han, Mesenchymal stem cells with Sirt1 overexpression suppress breast tumor growth via chemokine-dependent natural killer cells recruitment, Sci Rep 6: ((2016) ), 1–11. doi: 10.1038/srep35998. |
[40] | E. Rai, S. Sharma, S. Kaul, K. Jain, K. Matharoo, A.S. Bhanwer and R.N.K. Bamezai, The interactive effect of SIRT1 promoter region polymorphism on type 2 diabetes susceptibility in the north indian population, PLoS One 7: ((2012) ), 7–12. doi: 10.1371/journal.pone.0048621. |
[41] | S. Maeda, D. Koya, S.I. Araki, T. Babazono, T. Umezono, M. Toyoda, K. Kawai, M. Imanishi, T. Uzu, D. Suzuki, H. Maegawa, A. Kashiwagi, Y. Iwamoto and Y. Nakamura, Association between single nucleotide polymorphisms within genes encoding sirtuin families and diabetic nephropathy in Japanese subjects with type 2 diabetes, Clin Exp Nephrol 15: ((2011) ), 381–390. doi: 10.1007/s10157-011-0418-0. |
[42] | C.C.J.M. Simons, L.J. Schouten, R.W. Godschalk, F.J. van Schooten, P.A. van den Brandt and M.P. Weijenberg, Sirtuin 1 genetic variation, energy balance and colorectal cancer risk by sex and subsite in the Netherlands Cohort Study, Sci Rep 8: ((2018) ), 4–13. doi: 10.1038/s41598-018-34728-6. |
[43] | S.M. Rizk, N.N. Shahin and O.G. Shaker, Association between SIRT1 gene polymorphisms and breast cancer in Egyptians, PLoS One 11: ((2016) ), 1–19. doi: 10.1371/journal.pone.0151901. |
[44] | N. Kawaguchi-ihara, K. Arakawa and I.M. Id, The correlation of salivary telomere length and single nucleotide polymorphisms of the ADIPOQ , SIRT1 and FOXO3A genes with lifestyle-related diseases in a Japanese population, PLoS One 16: ((2021) ), 1–15. doi: 10.1371/journal.pone.0243745. |
[45] | Z. Shafieian, G. Bahari, M. Hashemi and A. Nakhaee, SIRT1 gene polymorphisms are associated with urinary bladder cancer in an Iranian population, Reports Biochem Mol Biol 8: ((2019) ), 194–199. |
[46] | L. Polito, P.G. Kehoe, G. Forloni and D. Albani, The molecular genetics of sirtuins: Association with human longevity and age-related diseases, Int J Mol Epidemiol Genet 1: ((2010) ), 214–225. |
[47] | W.G. Zhang, X.J. Bai and X.M. Chen, SIRT1 variants are associated with aging in a healthy Han Chinese population, Clin Chim Acta 411: ((2010) ), 1679–1683. doi: 10.1016/j.cca.2010.06.030. |
[48] | M. Kuningas, M. Putters, R.G.J. Westendorp, P.E. Slagboom and D. Van Heemst, SIRT1 gene, age-related diseases, and mortality: The Leiden 85-plus study, Journals Gerontol – Ser A Biol Sci Med Sci 62: ((2007) ), 960–965. doi: 10.1093/gerona/62.9.960. |
[49] | G. Balducci, Stefano, Sacchetti, Massimo, Haxhi, Jonida, Orlando, Giorgio, D’Errico, Valeria, Fallucca, Sara, Menini, Stefano, Pugliese, Candidate gene association study conditioning on individual ancestry in patients with type 2 diabetes and metabolic syndrome from Mexico City, Diabetes Metab Res Rev 32: ((2014) ), 13–23. doi: 10.1002/dmrr. |
[50] | N. Mohtavinejad, A. Nakhaee, H. Harati, J. Poodineh and M. Afzali, SIRT1 gene is associated with cardiovascular disease in the Iranian population, Egypt J Med Hum Genet 16: ((2015) ), 117–122. doi: 10.1016/j.ejmhg.2014.11.005. |
[51] | C.R. Consiglio, S. Juliana Da Silveira, O.A. Monticielo, R.M. Xavier, J.C.T. Brenol and J.A.B. Chies, SIRT1 promoter polymorphisms as clinical modifiers on systemic lupus erythematosus, Mol Biol Rep 41: ((2014) ), 4233–4239. doi: 10.1007/s11033-014-3294-3. |
[52] | W. Tang, Y. Chen, X. Fang, Y. Wang, W. Fan and C. Zhang, SIRT1 rs3758391 and major depressive disorder: New data and meta-analysis, Neurosci Bull 34: ((2018) ), 863–866. doi: 10.1007/s12264-018-0235-5. |
[53] | Y. Kan, P. Ge, X. Wang, G. Xiao and H. Zhao, SIRT1 rs3758391 polymorphism and risk of diffuse large B cell lymphoma in a Chinese population, Cancer Cell Int 18: ((2018) ), 1–8. doi: 10.1186/s12935-018-0659-z. |
[54] | O.G. Shaker, M.S. Wadie, R.M.M. Ali and A. Yosry, SIRT1 gene polymorphisms and its protein level in colorectal cancer, Gene Reports 7: ((2017) ), 164–168. doi: 10.1016/j.genrep.2017.04.005. |
[55] | Y. Lv, S. Lin and F. Peng, SIRT1 gene polymorphisms and risk of lung cancer, Cancer Manag Res 9: ((2017) ), 381–386. doi: 10.2147/CMAR.S142677. |
[56] | Y. Shimoyama, K. Suzuki, N. Hamajima and T. Niwa, Sirtuin 1 gene polymorphisms are associated with body fat and blood pressure in Japanese, Transl Res 157: ((2011) ), 339–347. doi: 10.1016/j.trsl.2011.02.004. |
[57] | A. V. Peeters, S. Beckers, A. Verrijken, I. Mertens, P. Roevens, P.J. Peeters, W. Hul and L.F. Gaal, Association of SIRT1 gene variation with visceral obesity, Hum Genet 124: ((2008) ), 431–436. doi: 10.1007/s00439-008-0567-8. |
[58] | M. Lee, S. Choi, Y. Lee and H.H. Oh, The gender association of the SIRT1 rs7895833 polymorphism with pediatric obesity: A 3-year panel study, J Nutrigenet Nutrigenomics 9: ((2017) ), 265–275. doi: 10.1159/000454713. |
[59] | X.-G. Yue, Z.-G. Yang, Y. Zhang, G.-J. Qin and F. Liu, Correlations between SIRT1 gene polymorphisms and diabetic kidney disease, R Soc Open Sci 5: ((2018) ), 171871. doi: 10.1098/rsos.171871. |
[60] | Y. Zhao, J. Wei, X. Hou, H. Liu, F. Guo, Y. Zhou, Y. Zhang, Y. Qu, J. Gu, Y. Zhou, X. Jia, G. Qin and L. Feng, SIRT1 rs10823108 and FOXO1 rs17446614 responsible for genetic susceptibility to diabetic nephropathy, Sci Rep 7: ((2017) ), 1–9. doi: 10.1038/s41598-017-10612-7. |
[61] | U. Kilic, O. Gok, U. Erenberk, M.R. Dundaroz, E. Torun, Y. Kucukardali, B. Elibol-Can, O. Uysal and T. Dundar, A remarkable age-related increase in SIRT1 protein expression against oxidative stress in elderly: SIRT1 gene variants and longevity in human, PLoS One 10: ((2015) ), 1–19. doi: 10.1371/journal.pone.0117954. |
[62] | G. Gatta, and et al, Prognoses and improvement for head and neck cancers diagnosed in Europe in early 2000s: The EUROCARE-5 population-based study, Eur J Cancer 51: ((2015) ), 2130–2143. doi: 10.1016/j.ejca.2015.07.043. |
[63] | G. Cadoni, L. Giraldi, L. Petrelli, M. Pandolfini, M. Giuliani, G. Paludetti, R. Pastorino, E. Leoncini, D. Arzani, G. Almadori and S. Boccia, Prognostic factors in head and neck cancer: A 10-year retrospective analysis in a single-institution in Italy, Acta Otorhinolaryngol Ital 37: ((2017) ), 458–466. doi: 10.14639/0392-100X-1246. |
[64] | X.M. Yu, Y. Liu, T. Jin, J. Liu, J. Wang, C. Ma and X.L. Pan, The expression of SIRT1 and DBC1 in laryngeal and hypopharyngeal carcinomas, PLoS One 8: ((2013) ), 1–8. doi: 10.1371/journal.pone.0066975. |




