Transcription factor KLF13 inhibits AKT activation and suppresses the growth of prostate carcinoma cells
Abstract
BACKGROUND:
Krüppel-like factor 13 (KLF13), a member of the KLF family, is involved in the development of immunological diseases and tumor progression. However, the expression patterns and potential functions of KLF13 in prostate carcinoma are still unknown. Here, we aimed to study the roles and mechanisms of KLF13 in prostate cancer.
METHODS:
The expression levels of KLF13 was detected by Immunohistochemistry in prostate tumor tissues and the paired non-tumor tissues. The effects of KLF13 up-regulation was tested by performing CCK8, cell colon formation, flow cytometric analysis and measurement of tumor proliferation in nude mice. Signaling pathway was analyzed by Western blot.
RESULTS:
The current study, for the first time, found that KLF13 was downregulated in prostate tumor tissues as compared to the paired non-tumor tissues. The overexpression of KLF13 dramatically inhibited cell proliferation and induced apoptosis by suppressing the AKT pathway in human prostate cancer cells. Moreover, the ectopic expression of KLF13 efficiently delayed the onset of PC3 xenografts and inhibited the tumor growth in vivo.
CONCLUSIONS:
KLF13 functions as a tumor suppressor protein in PCa, and the pharmacological activation of KLF13 might represent a potential approach for the treatment of prostate cancer.
1.Introduction
Prostate cancer (PCa) is the most common malignant carcinoma and the second leading cause of cancer-related deaths among males in the USA [1]; also its rate in developing countries, including China, is continually rising [2]. Although localized PCa can be regulated by the androgen deprivation therapy, a majority of the patients eventually relapse and become castration-resistant [3]. Thus, it would be significant to clarify the molecular mechanism underlying PCa progression and identify new therapeutic targets.
Krüppel-like factor 13 (KLF13), also known as RFLAT-1, BTEB3, and FKLF-2, is a critical member of the KLF family with essential functions in cell proliferation, migration, and differentiation [4, 5]. The KLF family has 17 members and can be divided into three distinct groups based on their biological functions and characteristics. KLF13, along with KLF9, 10, 11, 14, and 16, belongs to subgroup 3, whose members putatively exhibit a repressive function by interaction with Sin3A, a common transcriptional co-repressor [6]. Reportedly, KLF13 also regulates the life cycle of human papillomavirus in cervical keratinocytes [7] and controls the differentiation of adipocyte through PPAR
In the present study, we found that KLF13 expression is downregulated in PCa tissues, and the overexpression markedly inhibited the phosphorylation of AKT and induced apoptosis. Strikingly, the restoration of KLF13 caused a significant regression of established tumors in vivo. Therefore, this study aimed to deduce a novel mechanism underlying the role of KLF13 in PCa, and the upregulation of KLF13 might serve as a novel therapeutic strategy for specific cancer.
2.Materials and methods
2.1Prostate cancer cell lines and tissue specimens
Human prostate tissue microarray (HProA180PG03) used in this study was purchased from Shanghai Outdo Biotech (Shanghai, China). It contains 90 cases of normal prostate tissues, 73 cases of primary PCa tissues and 17 cases of metastatic prostate cancer tissues. Detailed information of this tissue microarray is available at http://www.superchip.com.cn/product/detail_340.aspx.
Human prostate cancer cell lines, DU145 and PC3, were obtained from Renji Hospital Clinical Stem Cell Research Center (Shanghai, China). Cell lines were maintained in DMEM medium (HyClone, Logan, UT) supplemented with 10% fetal bovine serum (FBS, HyClone, Logan, UT), penicillin (100 U/ml) and streptomycin (100 mg/ml) at 37
2.2Real-time PCR analysis
RNAs from cultured cells were isolated using the TRIzol reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions, followed by reverse transcription into cDNA using PrimeScript RT Master Kit (TaKaRa, Dalian, China). The qRT-PCR was performed using the SYBR Green PCR Master Mix (TaKaRa) on the ABI7300 System (Applied Biosystems, Foster City, CA, USA). The relative expression levels of the gene were calculated using the 2
2.3Western blotting
Cells were washed two times with pre-chilled phosphate-buffered saline (PBS) and collected with lysis buffer. The protein concentrations of the lysates were detected using the BCA assay. An equivalent of 50
2.4Immunohistochemistry
The tumor tissue biochip was deparaffinized with xylene and rehydrated with alcohol gradient. Then, it was retrieved in 1 mM EDTA buffer at pH 8.0 for 30 min, followed by blocking endogenous peroxidases. Subsequently, the tissue biochip was probed with anti-KLF13 antibody (1:200, LifeSpan) at 4
2.5Cell viability assay
Cell growth was assessed using Cell Counting Kit-8 (CCK8, Dojindo, Shanghai, China). A total of 1500 cells/well were seeded in 96-well plates and medium containing doxycycline (Sigma-Aldrich, 0.25 mg/mL) was added after 24 h. 10
2.6Colony formation assay
Cells were cultured in 6-well plates (1000/well) for 24 h, and inducers (doxycycline, 0.25 mg/mL) were added at 37
2.7Flow cytometric assay
The cells were collected, washed two times with PBS, re-suspended in binding buffer, and stained with Annexin V and 7-AAD (1
2.8Lentivirus infection
Cell transfections were performed using Lipofectamine 2000 reagent (Invitrogen, Carlsbad, USA) and HEK293T cells according to the manufacturer’s instructions. PC3 and DU145 cells were transduced with pTRIPZ-lenti-KLF13 or pTRIPZ-lenti-Vector (Open Biosystems, Huntsville, AL, USA); the stable transformants were isolated using puromycin (3
2.9Tumor xenograft model
Male BALB/c nude mice, 5-6-week-old (SLAC Laboratory, Shanghai) were injected subcutaneously with PC3/KLF13 and PC3/CON cells (5.0
2.10Statistical methods
Statistical analyses were performed using the GraphPad Prism v7.0 software (GraphPad Software Inc., La Jolla, CA, USA). Student’s t-test was used to compare the two groups, and unpaired two-sided Student’s t-test was used for other comparisons. Kaplan-Meier survival analysis analyzed the tumor-free survival. Data are presented as the mean
Figure 1.
KLF13 is downregulated in PCa tissues. (A) KLF13 expression in PCa tissues from TCGA prostate adenocarcinoma dataset (Normal,
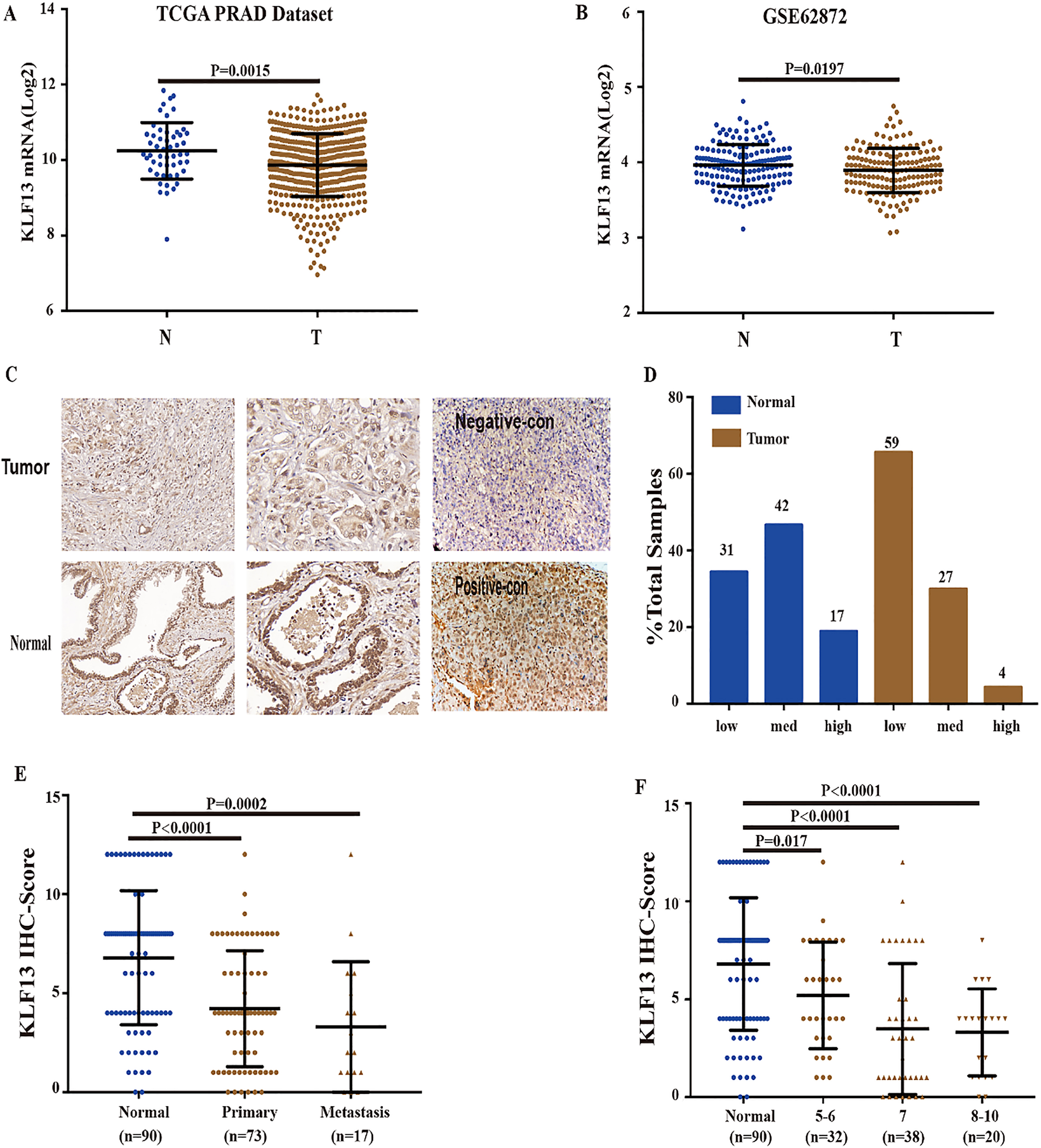
Figure 2.
Overexpression of KLF13 inhibits cell proliferation in vitro. (A and B) PC3 and DU145 cells transfected with pTRIPZ-KLF13 or pTRIPZ-vector were cultured with (Tet+) or without (Tet-) doxycycline (0.25 mg/mL) for 48 h. KLF13 expression was verified by Western blot. The viability of cells was assessed by CCK8 assay (OD value 450 nm) at the indicated time. (C and D) Colony formation rates of PC3 and DU145 cells treated with or without doxycycline for 10 days. Each value represents the mean
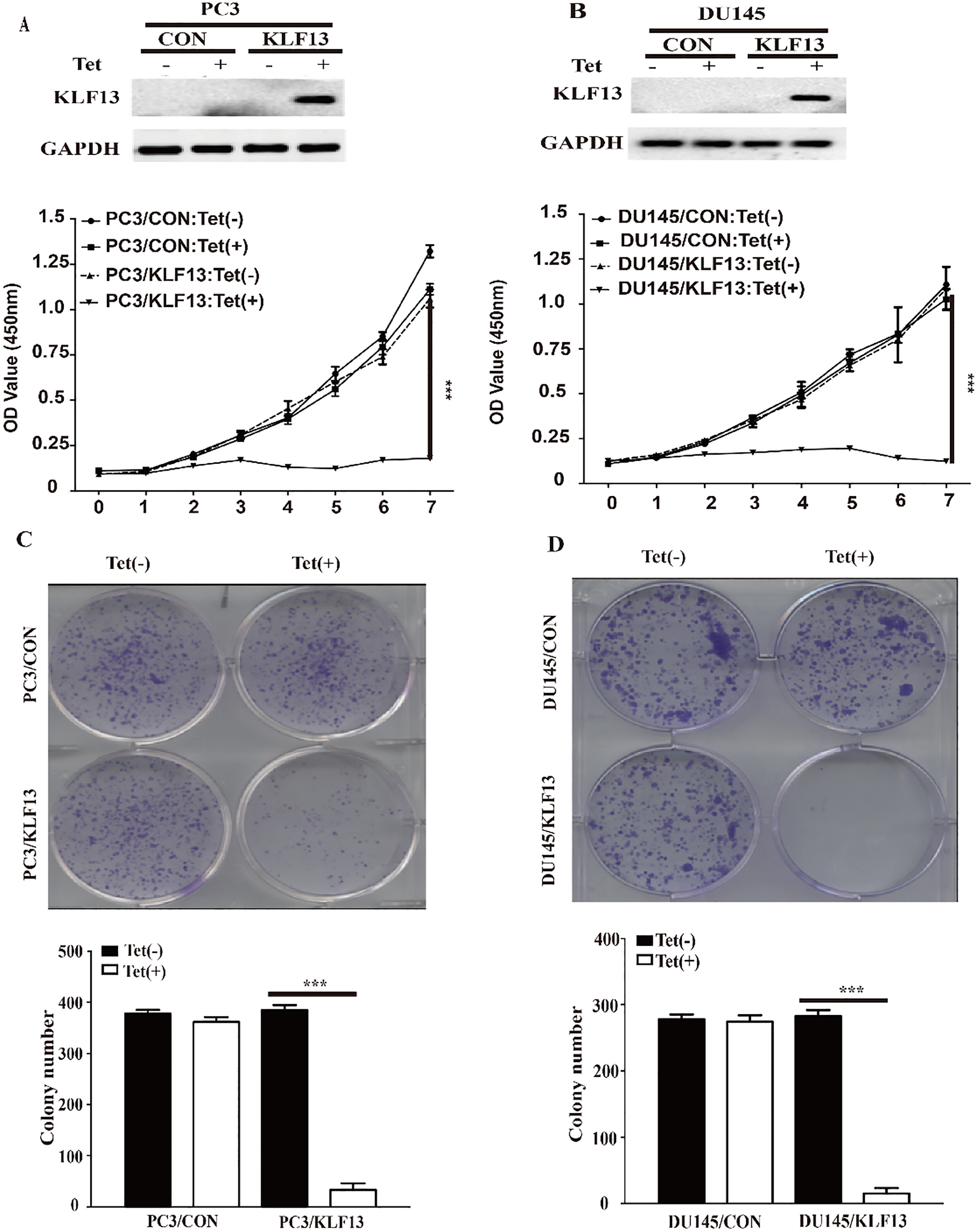
Figure 3.
KLF13 induces apoptosis and inhibits AKT activation. (A and B) Cells were cultured with or without doxycycline (0.25 mg/mL) and stained with AnnexinV/7-AAD. Apoptosis rates are shown on the left at 72 h. Statistical graphs for different groups at indicated time points are on the right. (C and D) The protein levels of PARP, BAX, P-AKT, AKT, P-S6, S6, and KLF13 were determined by Western blot after doxycycline treatment.
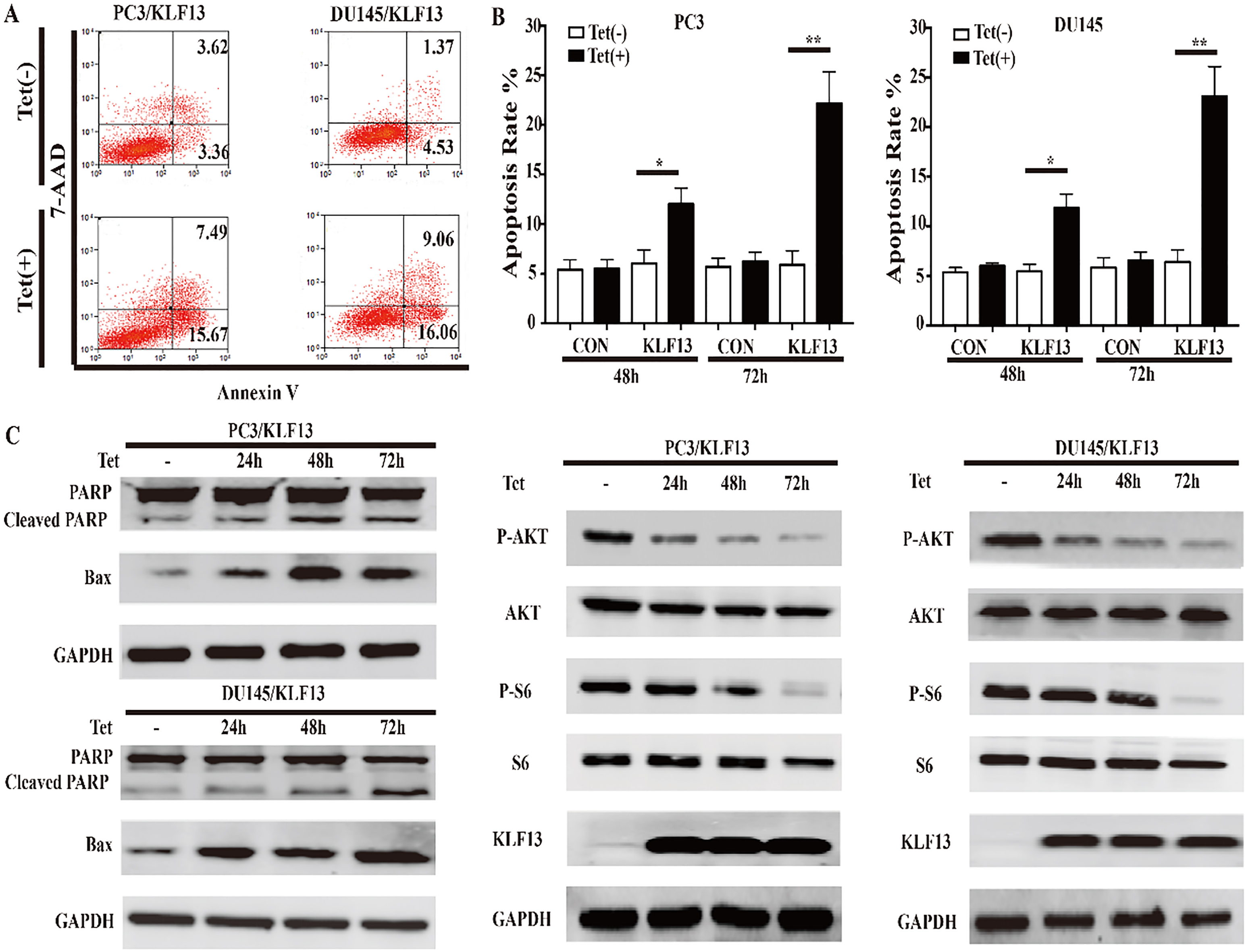
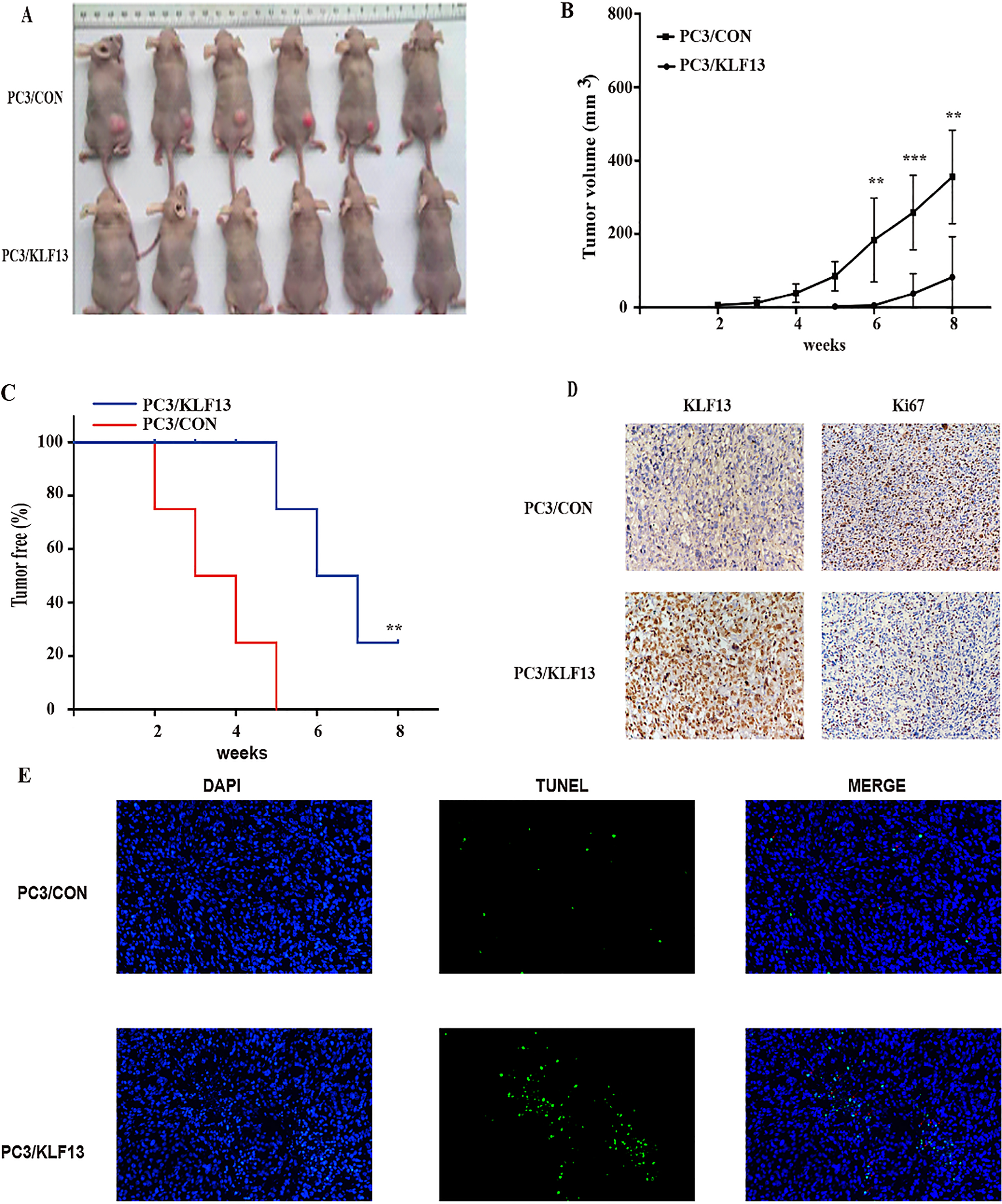
Figure 4.
KLF13 inhibits tumor growth of PCa in vivo. (A) The images of tumors in nude mice. (B) Tumor volume of PC3/KLF13 and PC3/CON cells. (C) Kaplan-Meier analysis of tumor onset. (D) IHC staining of KLF13 and Ki67 in tumor xenografts; original magnification 400
3.Results
3.1KLF13 is markedly downregulated in PCa tissues
Recent reports exhibited that KLF13 regulated the development of several human cancers; however, the biological functions in the development of prostate carcinoma are still unknown. To determine the expression levels of KLF13 in PCa and normal tissues, we firstly investigated its expression in two databases (GSE62827 and TCGA). As shown in Fig. 1A, the mRNA levels of KLF13 were markedly downregulated in PCa tissues as compared to adjacent non-tumor tissues. Notably, the analysis of 160 PCa cases and their paired adjacent non-tumor tissues in the Gene Expression Omnibus (GEO) dataset (GSE62827) revealed a consistent downregulation of KLF13 (Fig. 1B). To further validate these results, we performed IHC staining of KLF13 in 90 pairs of human prostate tissues. IHC analysis further revealed a decreased expression of KLF13 in PCa tissues (Fig. 1C and D) and we use tumor xenografts from PC3/CON and PC3/KLF13 cells as our negative and positive control of IHC staining. And the levels of KLF13 staining were significantly decreased in patients with metastatic PCa or high Gleason score (Fig. 1E and F). Collectively, these results indicated that KLF13 might be a tumor suppressor gene in PCa.
3.2Overexpression of KLF13 inhibits cell proliferation in vitro
The downregulation of KLF13 in PCa tissues suggested a suppression function in PCa progression. Therefore, we explored the effects of ectopic expression of KLF13 using CCK8 and colony formation assays. The expression of the KLF13 protein was effectively induced in the presence of doxycycline (Tet-on) (Fig. 2A and B). Strikingly, the CCK8 assay showed that the proliferation was much slower in KLF13-expressing (Tet+) cells as compared to control groups (Tet-) on day 7 (Fig. 2A and B). Subsequently, the colony formation assay showed that KLF13 significantly decreased the cell clone formation ability as compared to the control groups (Fig. 2C and D). These results indicated that the upregulation of KLF13 in PCa cells inhibits the cell proliferation ability.
3.3KLF13 induces apoptosis and inhibits AKT activation in PCa cells
Next, we used flow cytometry to examine whether KLF13 could induce cell apoptosis, and found that the overexpression of KLF13 remarkably induced apoptosis in PC3 and DU145 cells (Fig. 3A and B). Consistent with the above results, the protein levels of the cleaved PARP and BAX were increased significantly (Fig. 3C). The AKT pathway was highly activated in prostate cancer, and it contributed to tumor development and progression [19, 20]. To further understand the function of KLF13 in cell proliferation and apoptosis, we assessed the expression of several proteins involved in the AKT pathway by Western blot and found that the phosphorylation of Ser473 on AKT was inhibited by KLF13 overexpression. Furthermore, the level of phosphorylated S6 protein, the direct target of active AKT, was also decreased. These findings suggested that the overexpression of KLF13 induces apoptosis and inhibits AKT activation in PCa cells.
3.4KLF13 inhibits the growth of PC3 tumor xenografts
Since the expression of KLF13 inhibits the proliferation of PCa cells, we next examined this effect using tumor xenograft models. As shown in Fig. 4A and B, the overexpression of KLF13 significantly reduced the tumor size and suppressed the tumor growth. Moreover, the overexpression of KLF13 also delayed the tumor onset and improved tumor-free survival rates of mice (Fig. 4C). Consistent with the above observations, IHC staining of Ki67 revealed that the overexpression of KLF13 suppressed cell proliferation in vivo (Fig. 4D) and TUNEL assay showed that overexpression of KLF13 also induced apopotosis in vivo (Fig. 4E). These findings indicated that KLF13 efficiently inhibits tumor growth and tumorigenicity in vivo.
4.Discussion
The recent evidence stated that several KLFs function as tumor suppressor genes in PCa. For example, KLF4 is downregulated in PCa tissues, and the overexpression of KLF4 inhibits cell viability, clonogenicity, invasion, and migration in PC3 cells [15]. In PTEN-deficient mouse prostate tumors, KLF5 deletion can enhance tumor angiogenesis by attenuating PI3K/AKT signaling [11]. Moreover, KLF9 contributes to flutamide-induced apoptosis in LNcap cells [14]. Emerging studies have revealed that KLF13 plays a major role in regulating tumor cell differentiation and survival [7, 9, 10]. However, whether KLF13 acts as an oncogene or tumor suppressor is yet controversial. Reportedly, KLF13 acts as an oncogene in oral and cervical cancers [7, 21], whereas it inhibits cellular growth, suppresses transformation, and increases apoptosis in pancreatic cancer cells and mediates BCL2 repression in pediatric acute lymphoblastic leukemia [9, 10]. However, the physiological function and clinical significance of KLF13 in PCa have not yet been investigated. In this study, for the first time, we demonstrated that KLF13 is moderately downregulated in PCa tissues, and its expression was much lower in metastatic or high-grade PCa tissues. Intriguingly, the exogenous KLF13 expression suppressed the cell proliferation in vivo and in vitro and induced apoptosis in human PCa cells. These results strongly indicated that KLF13 might act as a tumor suppressor gene in PCa.
The AKT pathway is a significant contributor towards the progression of PCa [22] and is responsible for alterations in 42% of primary PCas and 100% of metastatic tumors [23]. The phosphorylation of AKT at Ser473 is a useful biomarker and correlated with a high rate of disease recurrence [24]. The activation of AKT in LNCaP diminishes the expression of p27 and accelerates the growth prostate tumor in LNcap xenografts [25]. Moreover, KLF5 and KLF9 have been reported to suppress the proliferation of PCa by affecting the AKT signaling [11, 14]. CYP1A1 could upregulate the protein expression of AKT in breast cancer [26] and KLF13 binds the CYP1A1 promoter and suppress the expression of CYP1A, so we guess that KLF13 may suppress the activation of AKT [27]. In this study, we observed that the overexpression of KLF13 suppressed the AKT signaling pathway in PC3 and DU145 cells, while the decreased phosphorylation of AKT might contribute to cell apoptosis. However, the molecular mechanism underlying KLF13 in the progression of PCa necessitates further investigations.
In summary, the current study revealed that KLF13 is downregulated in PCa tissues, exerting a suppressive function on the development of PCa. Therefore, understanding the precise role of KLF13 in AKT signaling and the pathogenesis of PCa would provide useful therapeutic strategies against the progression of this disease.
Acknowledgments
The Shanghai Key Medical Specialty (Program, Grant number: ZK2015B04) and Shanghai Municipal Commission of Health and Family Planning (Program, Grant number: 201440493) supported this study.
Conflict of interest
The authors declare that they have no conflicts of interest.
References
[1] | R.L. Siegel, K.D. Miller and A. Jemal, Cancer Statistics, 2017, CA Cancer J Clin 67: ((2017) ), 7–30. |
[2] | W. Chen, R. Zheng, H. Zeng, S. Zhang and J. He, Annual report on status of cancer in China, 2011, Chin J Cancer Res 27: ((2015) ), 2–12. |
[3] | G. Attard, C. Parker, R.A. Eeles, F. Schröder, S.A. Tomlins, I. Tannock, C.G. Drake and J.S. de Bono, Prostate cancer, The Lancet 387: ((2016) ), 70–82. |
[4] | A. Song, Y.F. Chen, K. Thamatrakoln, T.A. Storm and A.M. Krensky, RFLAT-1: A new zinc finger transcription factor that activates RANTES gene expression in T lymphocytes, Immunity 10: ((1999) ), 93–103. |
[5] | M.P. Tetreault, Y. Yang and J.P. Katz, Kruppel-like factors in cancer, Nat Rev Cancer 13: ((2013) ), 701–713. |
[6] | J. Kaczynski, J.S. Zhang, V. Ellenrieder, A. Conley, T. Duenes, H. Kester, B. van Der Burg and R. Urrutia, The Sp1-like protein BTEB3 inhibits transcription via the basic transcription element box by interacting with mSin3A and HDAC-1 co-repressors and competing with Sp1, J Biol Chem 276: ((2001) ), 36749–36756. |
[7] | W. Zhang, S. Hong, K.P. Maniar, S. Cheng, C. Jie, A.W. Rademaker, A.M. Krensky and C. Clayberger, KLF13 regulates the differentiation-dependent human papillomavirus life cycle in keratinocytes through STAT5 and IL-8, Oncogene 35: ((2016) ), 5565–5575. |
[8] | S. Jiang, H. Wei, T. Song, Y. Yang, F. Zhang, Y. Zhou, J. Peng and S. Jiang, KLF13 promotes porcine adipocyte differentiation through PPARgamma activation, Cell Biosci 5: ((2015) ), 28. |
[9] | M.E. Fernandez-Zapico, G.A. Lomberk, S. Tsuji, C.J. DeMars, M.R. Bardsley, Y.H. Lin, L.L. Almada, J.J. Han, D. Mukhopadhyay, T. Ordog, N.S. Buttar and R. Urrutia, A functional family-wide screening of SP/KLF proteins identifies a subset of suppressors of KRAS-mediated cell growth, Biochem J 435: ((2011) ), 529–537. |
[10] | D.H. Jing, V.A. Bhadri, D. Beck, J.A.I. Thoms, N.A. Yakob, J.W.H. Wong, K. Knezevic, J.E. Pimanda and R.B. Lock, Opposing regulation of BIM and BCL2 controls glucocorticoid-induced apoptosis of pediatric acute lymphoblastic leukemia cells, Blood 125: ((2015) ), 273–283. |
[11] | X. Ci, C. Xing, B. Zhang, Z. Zhang, J.J. Ni, W. Zhou and J.T. Dong, KLF5 inhibits angiogenesis in PTEN-deficient prostate cancer by attenuating AKT activation and subsequent HIF1alpha accumulation, Mol Cancer 14: ((2015) ), 91. |
[12] | H.J. He, X.F. Gu, W.H. Xu, D.J. Yang, X.M. Wang and Y. Su, Kruppel-like factor 8 is a novel androgen receptor co-activator in human prostate cancer, Acta Pharmacol Sin 34: ((2013) ), 282–288. |
[13] | G. Narla, A. DiFeo, S. Yao, A. Banno, E. Hod, H.L. Reeves, R.F. Qiao, O. Camacho-Vanegas, A. Levine, A. Kirschenbaum, A.M. Chan, S.L. Friedman and J.A. Martignetti, Targeted inhibition of the KLF6 splice variant, KLF6 SV1, suppresses prostate cancer cell growth and spread, Cancer Res 65: ((2005) ), 5761–5768. |
[14] | P. Shen, J. Sun, G. Xu, L. Zhang, Z. Yang, S. Xia, Y. Wang, Y. Liu and G. Shi, KLF9, a transcription factor induced in flutamide-caused cell apoptosis, inhibits AKT activation and suppresses tumor growth of prostate cancer cells, Prostate 74: ((2014) ), 946–958. |
[15] | J. Wang, R.F. Place, V. Huang, X. Wang, E.J. Noonan, C.E. Magyar, J. Huang and L.C. Li, Prognostic value and function of KLF4 in prostate cancer: RNAa and vector-mediated overexpression identify KLF4 as an inhibitor of tumor cell growth and migration, Cancer Res 70: ((2010) ), 10182–10191. |
[16] | A. Bar-Shira, N. Matarasso, S. Rosner, D. Bercovich, H. Matzkin and A. Orr-Urtreger, Mutation screening and association study of the candidate prostate cancer susceptibility genes MSR1, PTEN, and KLF6, Prostate 66: ((2006) ), 1052–1060. |
[17] | G. Narla, K.E. Heath, H.L. Reeves, D. Li, L.E. Giono, A.C. Kimmelman, M.J. Glucksman, J. Narla, F.J. Eng, A.M. Chan, A.C. Ferrari, J.A. Martignetti and S.L. Friedman, KLF6, a Candidate Tumor Suppressor Gene Mutated in Prostate Cancer, Science, 294: ((2001) ), 2563–2566. |
[18] | E.H. Seppala, V. Autio, P. Duggal, T. Ikonen, U.H. Stenman, A. Auvinen, J.E. Bailey-Wilson, T.L. Tammela and J. Schleutker, KLF6 IVS1 -27G > A variant and the risk of prostate cancer in Finland, Eur Urol 52: ((2007) ), 1076–1081. |
[19] | C.W. Kinkade, M. Castillo-Martin, A. Puzio-Kuter, J. Yan, T.H. Foster, H. Gao, Y. Sun, X. Ouyang, W.L. Gerald, C. Cordon-Cardo and C. Abate-Shen, Targeting AKT/mTOR and ERK MAPK signaling inhibits hormone-refractory prostate cancer in a preclinical mouse model, Journal of Clinical Investigation ((2008) ). |
[20] | P.K. Majumder and W.R. Sellers, Akt-regulated pathways in prostate cancer, Oncogene 24: ((2005) ), 7465–7474. |
[21] | B.J. Henson and S.M. Gollin, Overexpression of KLF13 and FGFR3 in oral cancer cells, Cytogenet Genome Res 128: ((2010) ), 192–198. |
[22] | R.L. Bitting and A.J. Armstrong, Targeting the PI3K/Akt/mTOR pathway in castration-resistant prostate cancer, Endocr Relat Cancer 20: ((2013) ), R83–99. |
[23] | B.S. Taylor, N. Schultz, H. Hieronymus, A. Gopalan, Y. Xiao, B.S. Carver, V.K. Arora, P. Kaushik, E. Cerami, B. Reva, Y. Antipin, N. Mitsiades, T. Landers, I. Dolgalev, J.E. Major, M. Wilson, N.D. Socci, A.E. Lash, A. Heguy, J.A. Eastham, H.I. Scher, V.E. Reuter, P.T. Scardino, C. Sander, C.L. Sawyers and W.L. Gerald, Integrative genomic profiling of human prostate cancer, Cancer Cell 18: ((2010) ), 11–22. |
[24] | J.I. Kreisberg, S.N. Malik, T.J. Prihoda, R.G. Bedolla, D.A. Troyer, S. Kreisberg and P.M. Ghosh, Phosphorylation of Akt (Ser(473)) is an excellent predictor of poor clinical outcome in prostate cancer, Cancer Research 64: ((2004) ), 5232–5236. |
[25] | J.R. Graff, B.W. Konicek, A.M. McNulty, Z. Wang, K. Houck, S. Allen, J.D. Paul, A. Hbaiu, R.G. Goode, G.E. Sandusky, R.L. Vessella and B.L. Neubauer, Increased AKT activity contributes to prostate cancer progression by dramatically accelerating prostate tumor growth and diminishing p27Kip1 expression, J Biol Chem 275: ((2000) ), 24500–24505. |
[26] | A.G. Peres, R. Zamboni, I.L. King and J. Madrenas, Suppression of CYP1 members of the AHR response by pathogen-associated molecular patterns, Journal of Leukocyte Biology 102: ((2017) ), 1471–1480. |
[27] | J.A. Kaczynski, A.A. Conley, M. Fernandez Zapico, S.M. Delgado, J.S. Zhang and R. Urrutia, Functional analysis of basic transcription element (BTE)-binding protein (BTEB) 3 and BTEB4, a novel Sp1-like protein, reveals a subfamily of transcriptional repressors for the BTE site of the cytochrome P4501A1 gene promoter, Biochem J 366: ((2002) ), 873–882. |




