Mutations in Non-Tumoral Human Urothelium: Disease Prelude or Epilogue?
Abstract
Bladder cancer is characterized by high rates of recurrence and multifocality, features which have commonly been associated with the colonization of widespread areas of non-neoplastic urothelium by mutant cells, a phenomenon known as field change. Whether mutant fields in the bladder arise from tumor cells or develop from the accumulation of somatic mutations followed by clonal expansions of non-transformed progenitor cells during lifetime remains unanswered. In this issue, Strandgaard et al. perform a deep-sequencing analysis of paired samples of tumor and histologically normal-appearing urothelium from four patients with advanced bladder cancer. By using a careful validation process, they report several mutations exclusive of normal, non-neoplastic tissue, suggesting that multiple fields precede (or develop independently from) the disease. Here, we discuss the main results from this work and elaborate on the biological implications and open questions in the context of normal somatic clonal evolution and cancer risk. We finish providing some general guidelines for future experiments to resolve the role of field changes in bladder carcinogenesis and its possible clinical relevance.
Bladder cancer is a highly recurrent disease and frequently presents multifocally, as physically-independent synchronous tumors that often share a clonal origin [1]. These features suggest the existence of a field cancerization effect in the urinary bladder; that is, the presence of genetic alterations in one or various non-neoplastic areas of the tissue that might originate from tumor-derived cells (Fig. 1, left column) or, alternatively, precede the disease (Fig. 1, right column) [1, 2]. While the magnitude of this phenomenon in the urothelium from individuals without bladder cancer remains unclear, so does the extent to which these transformations can be ascribed to the accumulation of somatic mutations in embryonic development or during lifetime, either as a consequence of aging or caused by exposure to carcinogens, which are important questions to be addressed.
Fig. 1
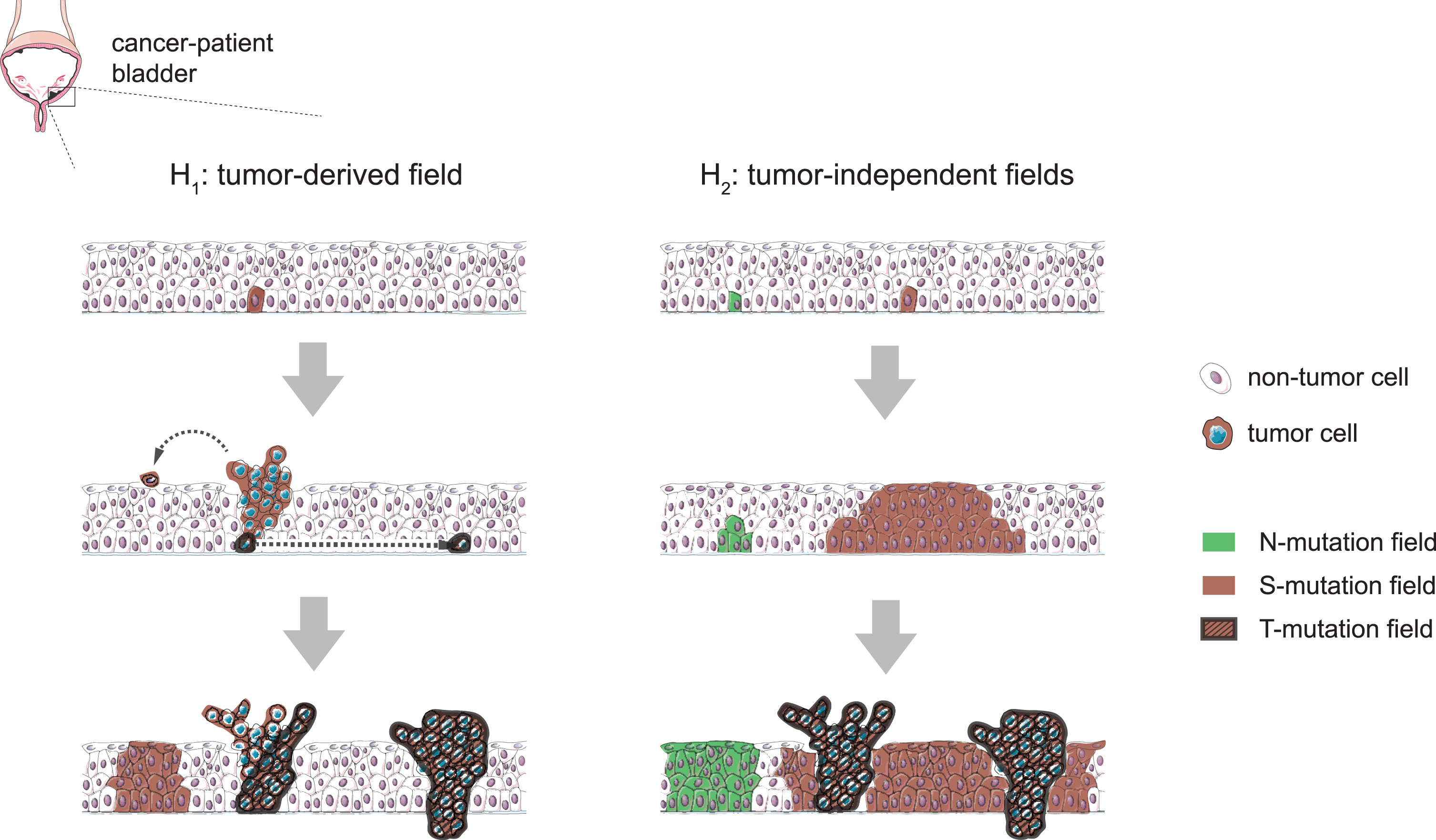
Hypothetical scenarios for the evolution of field changes in the bladder urothelium. Left: tumors develop from clones following the occasional mutation of cancer-driver genes and mutant fields extend as a consequence of tumor cell seeding (e.g. by shedding and implantation in distal regions or through horizontal trans-urothelial migration), preserving a clonal relationship with the tumor of origin. Right: mutant fields arise as a consequence of the accumulation of mutations in non-neoplastic proliferative cells and subsequent somatic clonal expansions. Cancer-driver mutations can occur within pre-established fields and cause tumors to arise as subclones of these, while other contemporary fields develop devoid of tumors and keep mutations not found in the tumors. This latter scenario is supported by N-mutations found in Strandgaard et al’s work, while we cannot discard a combination of the two models.
Next generation sequencing technologies are allowing the detection of mutations present in small areas of tissue at an unprecedented level of resolution, thus being suitable tools to characterize field changes in the urothelium. In this issue, Strandgaard et al. [3] perform deep-targeted amplicon sequencing of paired samples of tumor and histologically normal-appearing urothelium from four patients with advanced bladder cancer treated with radical cystectomy. The small cohort comprises two individuals with multifocal disease and two with unifocal disease. This study is a follow up of a previous one where the authors performed whole exome sequencing (WES) of the bulk tumors and subsequently analyzed genes found mutated in tumor across individual tumor biopsies and in laser microdissected (LMD) samples of non-neoplastic urothelium from each patient by deep-targeted amplicon sequencing [4]. Here, DNA from individual specimens is pooled and a large panel of 509 cancer-related genes is assayed in tumor and non-neoplastic sample pools from each patient. In this way, a more unbiased exploration can be achieved.
Three types of mutations are identified by Strandgaard and colleagues [3]: T-mutations, found exclusively in the tumor; S-mutations, found both in tumor and non-neoplastic urothelium; and N-mutations, found exclusively in non-neoplastic samples (hereafter referred to as normal tissue). The two former types are more abundant. However, still 29 out of the 206 point mutations detected are catalogued as N-mutations. These findings lead the authors to postulate the existence of multiple mutated fields in the adult bladder urothelium originating independently from tumors – possibly by the accumulation of mutations and somatic clonal expansions from non-neoplastic proliferative cells (Fig. 1). Interestingly, the two patients with multifocal disease present a higher prevalence of N-mutations, though larger cohorts would be needed to pinpoint the association of field effect with multifocality and recurrence.
The relatively modest number of mutations detected may also be regarded as a potential a priori limitation – perhaps discouraging, given the common incidence of background sequencing errors. However, the authors apply a stringent and meticulous filtering process. First, they use unique molecular identifiers (UMIs) for read collapsing to reduce error rates. Then, only variants found in at least 3 consensus reads with distinct UMIs are called (MuTect2). Putative N-mutations (initially, n = 224 in total) are subjected to read recounting in the tumor pool from the same patient, leading to reclassification of 118 of the initial calls as S-mutations. Finally, the remaining mutations are reevaluated in the previously generated WES data from bulk tumors and subjected to manual curation. As a consequence, the final set of 29 N-mutations represents high confidence mutations exclusive of non-tumoral tissue. The WES tumor data available and a digital droplet PCR test for a set of genes are used for further validation of the classification of N-, S- and T- mutations. In this regard, the authors are conservative and sacrifice the sensitivity to reliably differentiate low frequency mutations from sequencing errors and give answer to the main objective: to provide proof of concept evidence on the existence of (multiple) fields in the histologically normal bladder.
N-mutations are reported with lower variant allele fractions (VAFs) as compared to mutations in tumors. Yet, several N-mutations reach VAFs of ∼10%. Even though technical limitations preclude estimating clone sizes, this figure is consistent with wide mutated non-tumor fields. Considering that DNA was pooled from n = 6– 11 individual LMD normal samples per patient, it suggests that some of the N-mutants could have expanded to fully colonize one or more of the original samples. Intriguingly, annotations reveal no statistical difference in the predicted functional impact of N-mutations compared to S- or T-mutations, with all three categories being rich in high/moderate impact (nonsense and missense) mutations. This would support a scenario of somatic clonal expansions driven by positively-selected mutations that confer cells with growth advantage [5].
An important point that remains to be resolved is whether N-mutations are indeed associated with bladder carcinogenesis in these patients, or rather are independent manifestations of normal somatic evolution (e.g. aging, smoking) and do not necessarily promote tumorogenesis. Within the limited number of mutations called in this study, C > T substitutions at CpG context predominate among N-mutations, suggesting a signature of age-related mutagenesis, whereas APOBEC-related mutations prevail among S- and T-mutations [6]. Only 2 out of 29 N-mutations occurred in known bladder cancer drivers (BCOR, TBX3), despite the fact that more than half of the total N-sample hits are missense (15) and nonsense (2) mutations, considered of moderate and high impact to driver genes, respectively. This argues that field changes in normal-appearing urothelium may well represent a preneoplastic stage resulting from the accumulation of mutations related to age, lifestyle factors, or other exposures (the “prelude”). However, we cannot rule out that many of these mutations - even in cancer genes - may not necessarily herald significant increases in disease risk.
In the last few years, several publications have reported somatic mutations driving clonal expansions in normal tissues from healthy donors [7– 11]. Most of them have focused on high-renewal tissues (i.e. skin, esophageal epithelium, intestine and endometrium), with the exception of brain and liver. New sequencing studies involving healthy individuals are required to determine the extent of mutant colonization in a quiescent tissue such as normal bladder. The ideal design would combine histological imaging with ultra-deep sequencing of multiple synchronous bladder biopsies acquired from distinct sites of the same individual (i.e. from deceased organ donors). In addition, prospective longitudinal studies (e.g. biopsies obtained through cystoscopy) would provide highly informative temporal information. The benefits of such initiatives, when combined with larger cohorts of bladder cancer patients, could go beyond confirming that field effects originate in normal-appearing urothelium. As we elaborate a catalogue of mutations driving (or not) clonal expansions in normal tissue, it would be possible to refine the present list of exclusive N-mutations (e.g. some may actually be associated with the pathogenic condition or related to a history of carcinogen exposure). More importantly, we could disentangle which mutations shared with tumors are truly cancer drivers. It has recently been shown how some genes traditionally considered as cancer-related drivers appear more frequently mutated in normal aged human tissue than in the corresponding tumor types. Such is the case of NOTCH1 in the esophageal epithelium [8]. These findings inevitably urge to revise the catalogs of known cancer genes to filter out drivers of normal preneoplastic colonization that display no apparent role in malignant transformation and could even play a protective role [12]. One should emphasize that the formal notion of a clone as a group of cells sharing a common ancestry surpasses mutational connotations and that clone dynamics respond to mechanisms of progenitor cell proliferation and cell competition. “Clonal health” has been proposed as a homeostatic property of tissues that defines a healthy interaction among clonal populations inside the tissue [13]. Understanding the paradigms of proliferation and self-renewal and the forces that constrain mutant clone interactions in the urothelium could help to define clonal health status and how this property is lost in disease [14, 15].
The bladder lends itself particularly suitable to mutation monitoring through tests on urinary exfoliated cells and cell-free DNA [16]. In the near future, single-cell technologies may overcome the sensitivity of current bulk sequencing approaches once the methods of variant calling from single-cell DNA and/or RNA sequencing data mature. These can be promising tools for early cancer detection and to guide treatment once tumor-driving mutations are identified on a personalized basis. Cancer may originate from a complex patchwork of (preneoplastic) mutant clones in healthy tissue. Future studies aimed at understanding their evolutionary trajectories, following Strandgaard et al’s approach, will elucidate whether field effects allow monitoring bladder cancer risk and provide indications on the features of prospective tumors.
ACKNOWLEDGMENTS
The authors have no acknowledgments
FUNDING
GP is supported by a Talento program fellowship from Comunidad de Madrid. Work at the Epithelial Carcinogenesis Group is supported, in part, by a research grant from Fundación Científica de la Asociación Española Contra el Cáncer. CNIO is supported by Ministerio de Ciencia, Innovación y Universidades as a Centro de Excelencia Severo Ochoa SEV-2015-0510.
ETHICAL CONSIDERATIONS
This paper is a literature commentary and discussion that does not present any primary results of the study it describes. As such, it is exempt from any requirement for Institutional Review Board approval.
AUTHOR CONTRIBUTIONS
GP and FXR designed and supervised the work. GP produced the first draft. All authors contributed to write the final version.
CONFLICT OF INTEREST
FXR: Speaker Honoraria: Roche
Other Research Support: Boehringer-Ingelheim, VCN Biosciences
GP and LCF have no conflict of interest to report.
REFERENCES
[1] | Hafner C , Knuechel R , Stoehr R , Hartmann A . Clonality of multifocal urothelial carcinomas: 10 years of molecular genetic studies. Int J Cancer. (2002) ;101: :1–6. |
[2] | Slaughter DP , Southwick HW , Smejkal W . “Field cancerization” in oral stratified squamous epithelium. Clinical implications of multicentric origin. Cancer. (1953) ;6: :963–8. |
[3] | Strandgaard T , Nordentoft I , Lamy P , Christensen E , Thomsen MBH , Jensen JB , et al. Mutational analysis of field cancerization in bladder cancer. Bladder Cancer, in press. |
[4] | Thomsen MBH , Nordentoft I , Lamy P , Vang S , Reinert L , Mapendano CK , et al. Comprehensive multiregional analysis of molecular heterogeneity in bladder cancer. Sci Rep. (2017) ;7: :11702. |
[5] | Curtius K , Wright NA , Graham TA . An evolutionary perspective on field cancerization. Nat Rev Cancer. (2018) ;18: :19–32. |
[6] | Alexandrov LB , Jones PH , Wedge DC , Sale JE , Campbell PJ , Nik-Zainal S , et al. Clock-like mutational processes in human somatic cells. Nat Genet. (2015) ;47: :1402–7. |
[7] | Martincorena I , Roshan A , Gerstung M , Ellis P , Van Loo P , McLaren S , et al. Tumor evolution. High burden and pervasive positive selection of somatic mutations in normal human skin. Science. (2015) ;348: :880–6. |
[8] | Martincorena I , Fowler JC , Wabik A , Lawson ARJ , Abascal F , Hall MWJ , et al. Somatic mutant clones colonize the human esophagus with age. Science. (2018) ;362: :911–7. |
[9] | Moore L , Leongamornlert D , Coorens THH , Sanders MA , Ellis P , Dentro SC , et al. The mutational landscape of normal human endometrial epithelium. Nature. (2020) ;580: :640–6. |
[10] | Lodato MA , Rodin RE , Bohrson CL , Coulter ME , Barton AR , Kwon M , et al. Aging and neurodegeneration are associated with increased mutations in single human neurons. Science. (2018) ;359: :555–9. |
[11] | Blokzijl F , de Ligt J , Jager M , Sasselli V , Roerink S , Sasaki N , et al. Tissue-specific mutation accumulation in human adult stem cells during life. Nature. (2016) ;538: :260–4. |
[12] | Higa KC , DeGregori J . Decoy fitness peaks, tumor suppression, and aging. Aging Cell. (2019) ;18: :e12938. |
[13] | Fernández LC , Torres M , Real FX . Somatic mosaicism: on the road to cancer. Nat Rev Cancer. (2016) ;16: :43–55. |
[14] | Piedrafita G , Kostiou V , Wabik A , Colom B , Fernandez-Antoran D , Herms A , et al. A single-progenitor model as the unifying paradigm of epidermal and esophageal epithelial maintenance in mice. Nat Commun. (2020) ;11: :1429. |
[15] | Colom B , Alcolea MP , Piedrafita G , Hall MWJ , Wabik A , Dentro SC , et al. Spatial competition shapes the dynamic mutational landscape of normal esophageal epithelium. Nat Genet. (2020) ;52: :604–14. |
[16] | Patel KM , van der Vos KE , Smith CG , Mouliere F , Tsui D , Morris J , et al. Association Of Plasma And Urinary Mutant DNA With Clinical Outcomes In Muscle Invasive Bladder Cancer. Sci Rep. (2017) ;7: :5554. |




