Circulating Biomarkers in Bladder Cancer
Abstract
Bladder cancer is a molecularly heterogeneous disease characterized by multiple unmet needs in the realm of diagnosis, clinical staging, monitoring and therapy. There is an urgent need to develop precision medicine for advanced urothelial carcinoma. Given the difficulty of serial analyses of metastatic tumor tissue to identify resistance and new therapeutic targets, development of non-invasive monitoring using circulating molecular biomarkers is critically important. Although the development of circulating biomarkers for the management of bladder cancer is in its infancy and may currently suffer from lower sensitivity of detection, they have inherent advantages owing to non-invasiveness. Additionally, circulating molecular alterations may capture tumor heterogeneity without the sampling bias of tissue biopsy. This review describes the accumulating data to support further development of circulating biomarkers including circulating tumor cells, cell-free circulating tumor (ct)-DNA, RNA, micro-RNA and proteomics to improve the management of bladder cancer.
INTRODUCTION
Bladder cancer is the sixth most cancer in the united states and an estimated 75,000 new cases of urinary bladder cancer will be diagnosed in 2016 [1]. The median age of diagnosis is 73 years with two-thirds of cases occurring in men, making medical comorbidities a significant factor influencing patient management [2]. It lags behind other solid organ cancers with one of the lowest increases in 5-year survival in recent decades.
A majority of new cases are non-muscle invasive bladder cancer (NMIBC) with tumors largely confined to the mucosa (Ta-70%) or less often the submucosa (T1-20–25%) or flat high grade lesions/Carcinoma in situ (CIS-5–10%) [3]. The natural history of NMIBC is characterized by a tendency to recur locally as a function of stage, grade, size and multiplicity. Thus, NMIBC remains one of the few examples of a malignancy where serial tumor profiling separated by time is feasible due to repeated cystoscopic biopsies and resections [4].
For the treatment of muscle invasive bladder cancer (MIBC), neoadjuvant cisplatin based combination chemotherapy followed by radical cystectomy (RC) is proven to prolong survival [5, 6]. Adjuvant cisplatin-based combination therapy is considered reasonable in those with extra-vesical or node positive disease following RC, in those who have not received neoadjuvant therapy [7]. Maximal cystoscopic resection followed by concurrent chemoradiotherapy allows bladder preservation in selected patients based on tumor and patient characteristics [8]. However, long-term survival and cure remain elusive for most patients with extra-vesical or node positive disease at the time of RC. Moreover, clinical staging is suboptimal, with approximately 30–50% of cT2N0 patients upstaged at the time of RC [9].
The overall survival for advanced disease remains abysmal even with optimal first line cisplatin-based combination chemotherapy. The median overall survival (OS) is approximately 15 months, with 5-year OS rate of approximately 5% [10]. Moreover, the majority of patients do not receive cisplatin-based chemotherapy, partly due to renal dysfunction, poor performance status (PS) ≥ 2 or comorbidities (cardiac dysfunction, neuropathy, hearing loss) [11, 12]. Second-line and salvage systemic chemotherapy with taxanes or vinflunine yields even more dismal outcomes with a median OS of 6 to 8 months [13]. Recently, atezolizumab, a programmed death (PD)-ligand (L)-1 inhibitor, has been added to the therapeutic armamentarium for post-platinum therapy [14]. However, the median OS with atezolizumab remains around 8 months and overall response rate (ORR) is approximately 15%, although the excellent tolerability and duration of response represent a major advance for these patients.
All of the regimens above have been developed in unselected patients. There is an urgent need for newer agents to provide large increments of outcomes in appropriately selected patients, i.e. precision medicine. Given the difficulty of serial analyses of metastatic tumor tissue to study evolving mechanisms of resistance and new therapeutic targets, the development of non-invasively obtained circulating biomarkers for molecular tumor profiling is a priority. Moreover, early detection and clinical staging are suboptimal with currently employed modalities. Hence, there is an urgent need to identify superior non-invasive molecular tools for bladder cancer detection, staging, surveillance and therapy. This review will provide an overview of circulating biomarkers to enhance therapy for UC (Fig. 1).
CURRENTLY EMPLOYED CLINICAL AND LABORATORY PROGNOSTIC FACTORS
Risk stratification of patients based on traditional laboratory and clinical markers has been reported in several studies. The importance of stage, grade, multiplicity and prior recurrences is recognized in NMIBC [15]. Similarly, in the MIBC setting, pathologic stage confers a major prognostic impact [16]. In the first-line cisplatin-based chemotherapy setting, PS and visceral metastasis were identified to confer poor prognosis almost 2 decades ago [17]. Thereafter, a couple of other first-line models have further refined prognostication by the addition of other variables such as albumin, hemoglobin and leukocytosis [18, 19]. In the salvage therapy context, PS, hemoglobin and liver metastasis were described to be major adverse prognostic factors. This model has also been enhanced by the addition of treatment-free interval and albumin as additional prognostic factors [20, 21].
CURRENTLY APPROVED AND EMERGING NON-INVASIVE URINARY BIOMARKERS
While there are no commercially approved circulating molecular biomarkers for use in the clinic to manage bladder cancer, multiple urinary biomarkers have been commercially approved for NMIBC detection and surveillance using mostly protein based assays and a DNA based assay [22]: BTA Stat (Polymedco), BTA Trak (Polymedco), NMP22 Bladder Cancer Test (Matritech), NMP22 BladderChek (Matritech), ImmunoCyt/uCyt+ (DiagnoCure) and UroVysion (Abbott Molecular). The BTA (bladder tumor antigens) detect human complement factor H related protein produced by bladder cancer cells, with the BTA TRAK being a quantitative assay [23] while the BTA STAT is a qualitative test [24]. The NMP (Nuclear Matrix proteins) are proteins that play an important role in the structural framework of the nucleus and are involved in DNA replication and regulation of gene expression [25]. The NMP22 Bladder Cancer test kit is quantitative, while the NMP22 BladderChek test is a qualitative assay [26, 27]. The ImmunoCyt/uCyt+ - detects exfoliated bladder cancer cells in urine using fluorescent monoclonal antibodies for a high molecular weight form of carcinoembryonic antigen and 2 bladder tumor cell–associated mucins [28]. The UroVysion test uses a multi-targeted FISH assay to identify UC related chromosomal alterations- aneuploidy for chromosomes 3,7 and 17 and loss of 9p21 locus of the p16 tumor suppressor gene [29, 30]. Unfortunately, although many of these urinary markers are quite sensitive compared with cytology, they suffer from low specificity and cannot replace cystoscopy.
Other novel urinary assays with preliminary promise include DNA alteration assays in frequently mutated genes and protein assays (e.g. telomerase) [31, 32]. In one study, urinary cell free (cf)-DNA and cellular DNA was analyzed and compared to matched formalin-fixed paraffin embedded (FFPE) tumor DNA in 23 patients. Urinary DNA was highly representative of DNA derived from tumors. cfDNA from urine had a higher tumor genome border and allowed greater detection (90%) of key somatic mutations in (BRAF, KRAS, EGFR, IDH1, IDH2, PTEN, PIK3CA, NRAS and TP53) than cellular DNA from urine [32]. Additionally, mutations of telomerase reverse transcriptase (TERT) promoter have been detected in urinary cfDNA, which mirrors tumor tissue [31, 33–35].
TUMOR TISSUE ALTERATIONS TO GUIDE THERAPY FOR BLADDER CANCER
The cancer genome atlas (TCGA) network identified recurrent tumor tissue mutations in genes involved in cell-cycle regulation, chromatin regulation and kinase signaling pathways, including potential therapeutic targets in PI3K/AKT/MTOR, receptor tyrosine kinases and MAPK pathways [36]. Indeed, bladder cancer appears to demonstrate one of the highest somatic tumor mutation burdens [37]. TCGA and other groups have also identified multiple intrinsic subtypes based on gene expression profiling [38–41]. Emerging data indicate that the basal subtype may be more chemo-sensitive, the luminalsubtype may be responsive to FGFR, HER2, and HER3 inhibitors and the mesenchymal/claudin-low subtype may be responsive to T-cell checkpoint inhibitors [42]. Retrospective analysis of tumor tissue from a large phase II trial evaluating atezolizumab suggested that the expression of PD-L1, mutation burden and intrinsic subtype may be complementary and assist in the selection of patients for PD-1/PD-L1 inhibitors [43, 44]. Cisplatin-based neoadjuvant chemotherapy appears more active in patients with MIBC harboring somatic alterations of DNA repair genes (ATM, RB1, FANCC, ERBB2 and ERCC2), while the p53-like intrinsic subtype may be platinum-resistant [45–48]. Emerging data indicate that targeted agents may demonstrate substantial activity in selected populations with appropriate somatic alterations [49–52].
POTENTIAL CIRCULATING BIOMARKER PLATFORMS
Circulating tumor cells: Enumeration and profiling
Circulating tumor cells (CTCs) have long been the targets for ‘liquid biopsies’ and may provide insights into tumor tissue alterations. Conversely, CTCs are subject to the bias of selecting viable tumor cells capable of circulating in blood, and may not be representative of the entire malignancy. A major challenge has been to improve the sensitivity of CTC detection methods.
As CTCs are present at very low concentrations in peripheral blood, enrichment is required prior to detection to improve sensitivity. Enrichment is based on physical or biological properties that can discriminate CTCs from normal cells in circulation. Density-based separation is based on differential migration due to differences in density, while negative enrichment uses antibody mediated removal of hematopoietic cells and other non-cancer cells in circulation. Magnetic activated cell sorting uses magnetically labeled antibodies to capture CTCs, with EpCAM commonly used for positive enrichment and is currently FDA-approved. Cell size has been used to guide separation using a filtration-based approach to isolate epithelial cells, but is limited in application due to size variations within single populations. Separation based on magnetopheretic mobility can also be used for enrichment but is limited by the targets chosen. Following enrichment, multiple techniques exist for detection. Fluorescence assisted cell sorting (FACS) uses a flow cytometry based assay resulting in high purity and ability to accommodate high flow rates of up to 50,000 cells per second. Only a limited amount of cells can be analyzed due to its throughput design, and cell viability can also be reduced. Fiber-optic array scanning technology cytometry allows analysis of larger volumes of blood, eliminating the need for enrichment and improving cell viability. Microfluidic devices which utilize electric field gradients for cell sorting or dielectreopheresis can allow single cell isolation, which has significant advantages over other cell separation methods. ISET or isolation by size of epithelial tumor cells performs filtration-based enrichment based on larger tumor cell size, followed by immunohistochemical or cytological evaluation. The Adna test utilizes reverse transcription-polymerase chain reaction (RT-PCR) to detect CTCs having tumor-associated transcripts but false positives due to contaminating nucleic acid and loss of cell viability are limitations. The microfluidic platform utilized in CTC-Chip uses EpCAM-coated micro posts to isolate target CTCs, and is currently limited to CTCs that express EpCAM. CellSearch, which is FDA approved for CTC detection in peripheral blood, uses magnetic activated cell sorting with anti-EpCAM antibodies for enrichment followed by detection using positive selection by anti-cytokeratin and negative selection by anti-CD45 (lymphocyte common antigen) fluorescent dyes. Since CellSearch uses epithelial markers, non-epithelial phenotypes or cancer cells that have undergone epithelial-mesenchymal transition (EMT) are not detectable by CellSearch and other available CTC detection methods [53]. CTCs that have undergone EMT may be potentially identified by markers such as vimentin, N-cadherin and Twist [54].
CTCs were detected in circulating blood in 44% of 33 patients with metastatic UC when using the CellSearch platform. Higher numbers of CTCs were seen in patients with greater number of metastatic sites [55]. These findings were replicated in other studies demonstrating CTCs in approximately 50% of patients with metastatic UC [56, 57]. CTCs may be more prevalent in pre-treated patients as suggested by a study of 70 patients with platinum refractory UC, in whom CTCs by CellSearch were detectable in 66%, with Her2 positive CTCs in 3 patients [58].
However, CTCs by CellSearch were detected in only 21% of 43 patients prior to undergoing RC, and did not correlate with more advanced pathological extra-vesical or node positive disease [59]. Another larger study of 100 consecutive patients undergoingRC identified preoperative CTCs by CellSearch in 23% of patients. The presence of CTCs was associated with higher risks of recurrence and cancer specific and overall mortality [60]. This study also reported high concordance between the HER2 immunohistochemistry (IHC) status of CTCs and the fluorescent in situ hybridization (FISH) gene expression status of corresponding primary tumors (14 of 22 patients, 64%) and lymph node metastases (100%). Thirteen of 16 patients (81%) with HER2-negative primary tumors also had HER2-negative CTCs, but only 1 of 4 of patients with HER2-positive primary tumors also had HER2-positive CTCs. In addition, heterogeneous HER2 expression was occasionally observed among the identified CTCs.
As many as 20% of 102 patients with T1G3 NMIBC exhibited CTCs by CellSearch, which predicted decreased time to first recurrence and time to progression to muscle-invasive or metastatic disease [61]. Thus, CTCs may be useful even in NMIBC to identify those with high risk of recurrence who could potentially benefit from early systemic therapy. CTCs were found in 24 of 54 patients (44%) with T1G3 NMIBC in another study and 92% of CTCs expressed survivin [62]. Similarly, CTC positive patients exhibited shorter disease-free survival.
Preliminarily, CTC profiling of bladder cancer specific cells appears feasible for MUC7, epidermal growth factor receptor (EGFR), cytokeratin-20, survivin, folate receptor α ligand and uroplakin II by RT-PCR [62–68]. Another study using a novel IsoFlux™ System microfluidic collection device appeared to demonstrate increased sensitivity to capture CTCs compared with CellSearch, and next generation sequencing could be performed for a selected panel of genes [69]. Newer methodologies to isolate a larger number of CTCs continue to be studied, which may enhance the ability to more comprehensively perform molecular profiling in most patients [70–72]. One study simultaneously assessed 2 platforms - the ScreenCell Cyto platform which uses a size-selective enrichment method followed by central pathological review and the immunomagnetic Adna TestSelect kit to assess gene expression level of EPCAM and MUC1 by RT-PCR - in 3 cohorts of patients undergoing neoadjuvant therapy for MIBC, first line MVAC for metastatic disease and second line anti-TGFβ therapy. The combined strategy to identify CTCs showed promise in the ability to detect relapse with concordant results across bothplatforms [73].
A meta-analysis including studies evaluating all stages of disease demonstrated an overall low sensitivity of 35.1% for the detection of CTCs using mostly PCR and CellSearch assays, although higher stages more commonly displayed CTCs [74]. The technology to improve the detection and yield of true CTCs is still in its infancy and will continue to undergo refinement. Overall, while CTC positive patients appear more likely to have advanced disease, their low sensitivity across multiple studies and across stages appears to limit their utility as a screening or diagnostic tool. Potentially, the further development of platforms designed to identify CTCs that have undergone EMT may improve the utility of this circulating biomarker.
Circulating tumor DNA profiling
Cell free DNA consists of fragments of 120 to 200 base pairs and is present in the circulation and originates from both healthy cells and tumor cells. Circulating tumor (ct)-DNA consists of trace fractions of overall circulating DNA. Somatic mutations in ctDNA are widely representative of the underlying tumor genome and may provide better understanding of tumor heterogeneity, with less ‘sampling error’ of tissue biopsies and could be more representative of disseminated disease. ctDNA is detectable in >75% of patients with advanced solid malignancies, being identified in all patients with CTCs [75]. A recent retrospective study demonstrated that ctDNA exhibiting somatic DNA variants was detectable in plasma and urine of 12 patients using droplet digital PCR even in patients with non-invasive bladder cancer, with higher levels preceding disease progression [76]. Furthermore, ctDNA variants disappeared in disease-free patients.
Massive parallel NGS (Next generation sequencing) was employed to evaluate a panel of 70 potentially therapeutically actionable genes (Guardant 360, Guardant Health, Palo Alto, CA) for mutations, amplification, fusions and indels in a recent study of patients with advanced urothelial carcinoma of the bladder [77]. Among 143 patients with advanced bladder cancer, 127 (89%) had detectable ctDNA for profiling. These data suggest that bladder cancer may exhibit one of the highest prevalences of ctDNA. Alterations were frequently observed in p53, DNA repair genes (BRCA1/2) cell-cycle controlling genes (FGFR2/3, APC, CDKN2A) epigenetic modifying genes (ARID1A) and kinase genes (EGFR, HER2, PIK3CA, KRAS, RAF) which resembled alterations previously reported from tumor tissue by the TCGA. Serial profiling was also presented for a subset of patients, demonstrating the extinction of pre-existing alterations and the emergence of new alterations, which may be hypothesized to confer resistance and represent new therapeutic targets. While lack of concordant tissue testing is a limitation of this study, further studies are needed to confirm that ctDNA mirrors the aggregate of genomic alterations in contemporaneously obtained tumor tissue from various locations, which will provide a method of assessing tumor response and early identification of drug resistance especially in the era of targeted therapies.
Methylation of circulating cell-free DNA
Epigenetic modifications have been identified as consistent major somatic molecular alterations in bladder cancer, with DNA methylation occurring frequently. Thus, methylated DNA has been studied as an epigenetic tumor marker, with some hypermethylation signatures being tumor specific [78]. Circulating methylated genes have been identified in the plasma of 27 bladder cancer patients in a study, 21 of whom had NMIBC with the remaining having muscle invasive disease [79]. p14ARF promoter hypermethylation was associated with multicentric foci, larger tumors and relapse. Seventeen cases demonstrated accord between plasma and tumor DNA alterations, while 6 cases exhibited a new alteration in plasma. Although the concordance rate was 63%, perfect concordance is not expected due to various factors including sampling bias of tumor biopsies and presumably, DNA from all sites of tumors and germline DNA in the circulation. Analysis of cell free DNA in other small studies of patients with invasive or non-invasive bladder cancer revealed hypermethylation of cell-free serum DNA, especially promoters of specific genes such as APC, GSTP1, TIG1, DAPK, p16 and cadherin genes, and some of these aberrations were associated with aggressive clinicopathologic features and conferred poor prognosis [80–84]. These data accord with and resemble data derived from analysis of NMIBC tumor tissue [82, 85].
RNA and Micro (mi)-RNA profiling
Tumor tissue gene expression profiling of MIBC has led to the proposal of major subtypes of bladder cancer including basal, luminal and p53 subtypes. Potentially, these classifications can be proposed based on circulating mRNA for more advanced stages of disease [39, 40]. Peripheral blood mononuclear cells (PBMCs) have been subjected to gene expression (RNA) profiling in multiple conditions, although bladder cancer has not been evaluated by this assay. Recently, robust platforms such as NanoString have been employed to assess gene expression from circulating blood in other settings, which utilizes minute quantities of RNA, and may be worth exploring in bladder cancer patients too [86].
MicroRNAs (miRNA) are a class of small, conserved non protein-coding RNAs that regulate gene expression by controlling translation of target mRNAs. Indeed, the potential relevance miRNAs in driving bladder cancer progression has been demonstrated from studies of tumor tissue also. For example, one study of tumor tissue demonstrated that miR-21, which downregulates the p53 pathway was increased in MIBC. Conversely, NMIBC exhibited miRNA expression profiles that led to increased FGFR3 expression [87]. miR expression profiles were prognostic and may assist in developing precision medicine. For example, high tumor levels of miR-21 and miR-372 were associated with poor outcomes, and miR-203 were associated with better outcomes in separate studies in the context of cisplatin-based chemotherapy for patients with advanced UC or MIBC, respectively [88, 89].
miRNAs are better able to circulate without significant degradation in blood and withstand processing due to smaller size. Their expression can be measured by real time quantitative PCR or miRNA microarray platforms [90]. Studies of serum miRNA expression in bladder cancer have identified miRNA profiles that may distinguish NMIBC from MIBC or confer prognostic impact. In a recent large cohort, genome-wide miRNA analysis by sequencing was performed followed by RT-PCR on serum from 207 patients with MIBC, 285 with NMIBC and 193 controls [91]. This study reported that miR-486-3p and miR-103a-3p were associated with overall survival of patients with MIBC. In another report, these authors also identified an association of miR-152 with tumor recurrence of NMIBC, and the potential value of a 6 miR panel to diagnose bladder cancer [92]. Plasma miR-205 was upregulated in 89 bladder cancer patients compared to 56 healthy control subjects and in MIBC compared to NMIBC patients [93]. Plasma miR-497 and miR-663b were identified to be potentially promising circulating diagnostic biomarkers in another study, which included mostly NMIBC (70%) evaluated 56 bladder cancer and 60 control patient samples in the training phase and 109 bladder cancer and 115 controls in the validation phase [94].
Proteomics
Proteomic studies have been conducted on various blood derived samples for a longer time, but are plagued by the non-specific nature of protein molecules in blood. Serum levels of soluble E-cadherin, MMP2, MMP7, endostatin, TGF-β and uPA have all been reported to be associated with higher pathologic stage or prognostic for poorer clinical outcomes in different settings [95–99]. High circulating interleukin (IL)-8 levels at baseline were associated with poor outcomes in the setting of sunitinib or pazopanib [100, 101]. More sophisticated serum protein analysis using matrix-assisted laser desorption ionization (MALDI) time-of-flight (TOF) mass spectrometry (MS) have been evaluated more recently. In one study of serum samples from 105 patients with bladder cancer, 98 healthy controls, and 45 prostate cancer patients, MALDI-TOF-MS demonstrated potential diagnostic utility in identifying patients with bladder cancer [102]. Furthermore, nuclear magnetic resonance (NMR)-based metabolomics has identified differences of serum metabolic profiles between bladder cancer and control subjects [103]. Interestingly, serum samples from bladder cancer patients exhibited decreased levels of isoleucine, leucine, tyrosine, lactate, glycine and citrate, coupled with increased levels of lipids and glucose.
Current discovery driven proteomic studies are still unable to detect uncommon proteins at low concentrations of 1 ng/ml or less, which are more likely to be biomarkers. The large dynamic concentration range of upto 12 orders of magnitude of plasma proteins can mask the targeted plasma proteins present in lower quantities. Despite advancements in techniques including sample fractionation and protein depletion, sensitivity remains low. Moreover, the standardization and reproducibility of results is hindered by technological challenges, variations due to time of collection, age, sex and also genetic variations [104]. Rapid enzymatic degradation of some proteins requiring immediate sample processing and significant variations in some proteins based on fasting vs non-fasting states entail attention to details of uniform timing of sample collections. Therefore, the development of circulating proteomic biomarkers faces intrinsic challenges and needs to occur with significant thought.
CONCLUSION
The development of molecular biomarkers, particularly circulating biomarkers, for the management of bladder cancer is in its infancy. Circulating biomarkers have inherent logistical advantages owing to their non-invasiveness, but suffer from lower sensitivity of detection and moderate correlation with tumor tissue alterations. Conversely, circulating molecular alterations may capture tumor heterogeneity without the sampling bias of tissue biopsy.
Moreover, serial assessment of circulating biomarkers is readily accomplished, which can facilitate cancer screening, knowledge of tumor biology, identify minimal residual or occult disease, early detection of recurrence, track resistance mechanisms and inform therapeutic intervention. Indeed, one recently initiated basket trial, TAPUR (Targeted Agent and Profiling Utilization Registry) allows the enrollment of patients to one of multiple arms of targeted therapy based on molecular alteration detected by ctDNA profiling using a Clinical Laboratory Improvement Amendments (CLIA)-certified, College of American Pathologists (CAP)-accredited laboratory, which has registered the assay with the National Institutes of Health (NIH) Genetic Test Registry. For example, given the preliminary success of ctDNA profiling to identify alterations in kinase genes, epigenetic modulating genes and p53, UC patients could be enrolled in trials such as TAPUR based on accredited ctDNA profiling. Notably, one phase III trial could not detect a favorable impact of adjuvant cisplatin-based chemotherapy following cystectomy in patients with tumor tissue p53 alteration by IHC, and this paradigm could be employed in metastatic disease using ctDNA profiling [105]. Circulating panels could be developed in conjunction with developing tumor tissue based profiling using the neoadjuvant therapy paradigm such as the randomized phase II neoadjuvant trial that compares GC vs. MVAC (NCT02177695) and evaluates COXEN (Coexpression Extrapolation) molecular tumor profiling, using an algorithm based on resistance patterns from a panel of 60 diverse cancer cell lines to predict sensitivity to chemotherapy [106].
Potentially, capture of data for multiple circulating biomarkers (CTCs, DNA including methylation status, RNA, miRNA, proteins) on multiplex platforms may be complementary and optimize a comprehensive circulating biomarker panel to advance the ultimate goals of early diagnosis, optimal clinicalstaging, monitoring and precision medicine. Moreover, the integration of molecular panels from interrogating another non-invasively obtained fluid, urine, may also complement circulating biomarker panels. Emerging imaging technology using novel radiotracers are also worthy of attention in this context, given their non-invasiveness. The task of studying and validating such comprehensive platforms in large prospective studies is undeniably challenging and will require a multidisciplinary and international effort in partnership with regulatory agencies.
RELEVANT DISCLOSURES
Guru Sonpavde: Consultant for Bayer, Sanofi, Pfizer, Novartis, Eisai, Janssen, Amgen, Astrazeneca, Merck, Genentech, Argos, Agensys; Research support to institution from Bayer, Onyx, Celgene, Boehringer-Ingelheim, Merck, Pfizer; Author for Uptodate; Speaker for Clinical Care Options
ACKNOWLEDGMENTS
None.
REFERENCES
[1] | Siegel RL , Miller KD , Jemal A . Cancer statistics, 2016. CA Cancer J Clin (2016) ;66: (1):7–30. |
[2] | Howlader N, Noone AM, Krapcho M, Miller D, Bishop K, Altekruse SF, Kosary CL, Yu M, Ruhl J, Tatalovich Z, Mariotto A, Lewis DR, Chen HS, Feuer EJ, Cronin KA (eds). SEER Cancer Statistics Review, 1975–2013, National Cancer Institute. Bethesda, MD, http://seer.cancer.gov/csr/1975_2013/, based on November 2015 SEER data submission, posted to the SEER web site, April 2016. |
[3] | Pasin E , Josephson DY , Mitra AP , Cote RJ , Stein JP . Superficial bladder cancer: An update on etiology, molecular development, classification, and natural history. Rev Urol (2008) ;10: (1):31–43. |
[4] | Sylvester RJ , van der Meijden AP , Oosterlinck W , Witjes JA , Bouffioux C , Denis L , et al. Predicting recurrence and progression in individual patients with stage Ta T1 bladder cancer using EORTC risk tables: A combined analysis of 2596 patients from seven EORTC trials. Eur Urol (2006) ;49: (3):466–5; discussion 75-7. |
[5] | Grossman HB , Natale RB , Tangen CM , Speights VO , Vogelzang NJ , Trump DL , et al. Neoadjuvant chemotherapy plus cystectomy compared with cystectomy alone for locally advanced bladder cancer. The New England Journal of Medicine (2003) ;349: (9):859–66. |
[6] | Griffiths G , Hall R , Sylvester R , Raghavan D , Parmar MK . International phase III trial assessing neoadjuvant cisplatin, methotrexate, and vinblastine chemotherapy for muscle-invasive bladder cancer: Long-term results of the BA06 30894 trial. J Clin Oncol (2011) ;29: (16):2171–7. |
[7] | Galsky MD , Stensland KD , Moshier E , Sfakianos JP , McBride RB , Tsao CK , et al. Effectiveness of Adjuvant Chemotherapy for Locally Advanced Bladder Cancer. J Clin Oncol (2016) ;34: (8):825–32. |
[8] | James ND , Hussain SA , Hall E , Jenkins P , Tremlett J , Rawlings C , et al. Radiotherapy with or without chemo-therapy in muscle-invasive bladder cancer. N Engl J Med (2012) ;366: (16):1477–88. |
[9] | Shariat SF , Palapattu GS , Karakiewicz PI , Rogers CG , Vazina A , Bastian PJ , et al. Discrepancy between clinical and pathologic stage: Impact on prognosis after radical cystectomy. Eur Urol (2007) ;51: (1):137–49; discussion 49-51. |
[10] | von der Maase H , Sengelov L , Roberts JT , Ricci S , Dogliotti L , Oliver T , et al. Long-term survival results of a randomized trial comparing gemcitabine plus cisplatin, with methotrexate, vinblastine, doxorubicin, plus cisplatin in patients with bladder cancer. J Clin Oncol (2005) ;23: (21):4602–8. |
[11] | Galsky MD , Hahn NM , Rosenberg J , Sonpavde G , Hutson T , Oh WK , et al. Treatment of patients with metastatic urothelial cancer “unfit” for Cisplatin-based chemotherapy. J Clin Oncol 29: (17):2432–8. |
[12] | De Santis M , Bellmunt J , Mead G , Kerst JM , Leahy M , Maroto P , et al. Randomized phase II/III trial assessing gemcitabine/carboplatin and methotrexate/carboplatin/vinblastine in patients with advanced urothelial cancer who are unfit for cisplatin-based chemotherapy: EORTC study 30986. J Clin Oncol (2012) ;30: (2):191–9. |
[13] | Sonpavde G , Sternberg CN , Rosenberg JE , Hahn NM , Galsky MD , Vogelzang NJ . Second-line systemic therapy and emerging drugs for metastatic transitional-cell carcinoma of the urothelium. The Lancet Oncology (2010) ;11: (9):861–70. |
[14] | Rosenberg JE , Hoffman-Censits J , Powles T , van der Heijden MS , Balar AV , Necchi A , et al. Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: A single-arm, multicentre, phase 2 trial. Lancet (2016) ;387: (10031):1909–20. |
[15] | Cambier S , Sylvester RJ , Collette L , Gontero P , Brausi MA , van Andel G , et al. EORTC Nomograms and Risk Groups for Predicting Recurrence, Progression, and Disease-specific and Overall Survival in Non-Muscle-invasive Stage Ta-T1 Urothelial Bladder Cancer Patients Treated with 1-3 Years of Maintenance Bacillus Calmette-Guerin. Eur Urol (2016) ;69: (1):60–9. |
[16] | Shariat SF , Karakiewicz PI , Palapattu GS , Amiel GE , Lotan Y , Rogers CG , et al. Nomograms provide improved accuracy for predicting survival after radical cystectomy. Clin Cancer Res (2006) ;12: (22):6663–76. |
[17] | Bajorin DF , Dodd PM , Mazumdar M , Fazzari M , McCaffrey JA , Scher HI , et al. Long-term survival in metastatic transitional-cell carcinoma and prognostic factors predicting outcome of therapy. J Clin Oncol (1999) ;17: (10):3173–81. |
[18] | Apolo AB , Ostrovnaya I , Halabi S , Iasonos A , Philips GK , Rosenberg JE , et al. Prognostic model for predicting survival of patients with metastatic urothelial cancer treated with cisplatin-based chemotherapy. Journal of the National Cancer Institute (2013) ;105: (7):499–503. |
[19] | Galsky MD , Moshier E , Krege S , Lin CC , Hahn N , Ecke T , et al. Nomogram for predicting survival in patients with unresectable and/or metastatic urothelial cancer who are treated with cisplatin-based chemotherapy. Cancer (2013) ;119: (16):3012–9. |
[20] | Sonpavde G , Pond GR , Fougeray R , Choueiri TK , Qu AQ , Vaughn DJ , et al. Time from prior chemotherapy enhances prognostic risk grouping in the second-line setting of advanced urothelial carcinoma: A retrospective analysis of pooled, prospective phase 2 trials. Eur Urol (2013) ;63: (4):717–23. |
[21] | Sonpavde G , Pond GR , Rosenberg JE , Bajorin DF , Choueiri TK , Necchi A , et al. Improved 5-Factor Prognostic Classification of Patients Receiving Salvage Systemic Therapy for Advanced Urothelial Carcinoma. J Urol (2016) ;195: (2):277–82. |
[22] | Mbeutcha A , Lucca I , Mathieu R , Lotan Y , Shariat SF . Current Status of Urinary Biomarkers for Detection and Surveillance of Bladder Cancer. Urol Clin North Am (2016) ;43: (1):47–62. |
[23] | Malkowicz SB . The application of human complement factor H-related protein (BTA TRAK) in monitoring patients with bladder cancer. Urol Clin North Am (2000) ;27: (1):63–73, ix. |
[24] | Raitanen MP , Marttila T , Nurmi M , Ala-Opas M , Nieminen P , Aine R , et al. Human complement factor H related protein test for monitoring bladder cancer. J Urol (2001) ;165: (2):374–7. |
[25] | He D , Zeng C , Brinkley BR . Nuclear matrix proteins as structural and functional components of the mitotic apparatus. Int Rev Cytol (1995) ;162B: :1–74. |
[26] | Su CK , Yang CR , Horng YY , Kao YL , Ho HC , Ou YC , et al. NMP22 in transitional cell carcinoma of the urinary bladder. J Chin Med Assoc (2003) ;66: (5):294–8. |
[27] | Grossman HB , Soloway M , Messing E , Katz G , Stein B , Kassabian V , et al. Surveillance for recurrent bladder cancer using a point-of-care proteomic assay. JAMA (2006) ;295: (3):299–305. |
[28] | Mian C , Pycha A , Wiener H , Haitel A , Lodde M , Marberger M . Immunocyt: A new tool for detecting transitional cell cancer of the urinary tract. J Urol (1999) ;161: (5):1486–9. |
[29] | Lotan Y , Bensalah K , Ruddell T , Shariat SF , Sagalowsky AI , Ashfaq R . Prospective evaluation of the clinical usefulness of reflex fluorescence in situ hybridization assay in patients with atypical cytology for the detection of urothelial carcinoma of the bladder. J Urol (2008) ;179: (6):2164–9. |
[30] | Skacel M , Fahmy M , Brainard JA , Pettay JD , Biscotti CV , Liou LS , et al. Multitarget fluorescence in situ hybridization assay detects transitional cell carcinoma in the majority of patients with bladder cancer and atypical or negative urine cytology. J Urol (2003) ;169: (6):2101–5. |
[31] | Fedriga R , Gunelli R , Nanni O , Bacci F , Amadori D , Calistri D . Telomerase activity detected by quantitative assay in bladder carcinoma and exfoliated cells in urine. Neoplasia (2001) ;3: (5):446–50. |
[32] | Togneri FS , Ward DG , Foster JM , Devall AJ , Wojtowicz P , Alyas S , et al. Genomic complexity of urothelial bladder cancer revealed in urinary cfDNA. Eur J Hum Genet (2016) ;24: (8):1167–74. |
[33] | Kinde I , Munari E , Faraj SF , Hruban RH , Schoenberg M , Bivalacqua T , et al. TERT promoter mutations occur early in urothelial neoplasia and are biomarkers of early disease and disease recurrence in urine. Cancer Res (2013) ;73: (24):7162–7. |
[34] | Allory Y , Beukers W , Sagrera A , Flandez M , Marques M , Marquez M , et al. Telomerase reverse transcriptase promoter mutations in bladder cancer: High frequency across stages, detection in urine, and lack of association with outcome. Eur Urol (2014) ;65: (2):360–6. |
[35] | Li C , Wu S , Wang H , Bi X , Yang Z , Du Y , et al. The C228T mutation of TERT promoter frequently occurs in bladder cancer stem cells and contributes to tumorigenesis of bladder cancer. Oncotarget (2015) ;6: (23):19542–51. |
[36] | Cancer Genome Atlas Research N. Comprehensive molecular characterization of urothelial bladder carcinoma. Nature (2014) ;507: (7492):315–22. |
[37] | Lawrence MS , Stojanov P , Polak P , Kryukov GV , Cibulskis K , Sivachenko A , et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature (2013) ;499: (7457):214–8. |
[38] | Comprehensive molecular characterization of urothelial bladder carcinoma. Nature (2014) ;507: (7492):315–22. |
[39] | Choi W , Porten S , Kim S , Willis D , Plimack ER , Hoffman-Censits J , et al. Identification of distinct Basal and luminal subtypes of muscle-invasive bladder cancer with different sensitivities to frontline chemotherapy. Cancer Cell (2014) ;25: (2):152–65. |
[40] | Damrauer JS , Hoadley KA , Chism DD , Fan C , Tiganelli CJ , Wobker SE , et al. Intrinsic subtypes of high-grade bladder cancer reflect the hallmarks of breast cancer biology. Proceedings of the National Academy of Sciences of the United States of America (2014) ;111: (8):3110–5. |
[41] | Sjodahl G , Lauss M , Lovgren K , Chebil G , Gudjonsson S , Veerla S , et al. A molecular taxonomy for urothelial carcinoma. Clin Cancer Res (2012) ;18: (12):3377–86. |
[42] | Hurst CD , Knowles MA . Molecular subtyping of invasive bladder cancer: Time to divide and rule? Cancer Cell (2014) ;25: (2):135–6. |
[43] | Ribas A , Tumeh PC . The future of cancer therapy: Selecting patients likely to respond to PD1/L1 blockade. Clin Cancer Res (2014) ;20: (19):4982–4. |
[44] | Rosenberg JE PD , Van Der Heijden MS , et al. PD-L1 expression, Cancer Genome Atlas (TCGA) subtype, and mutational load as independent predictors of response to atezolizumab (atezo) in metastatic urothelial carcinoma (mUC; IMvigor210). J Clin Oncol (2016) ;34: (suppl; abstr 104). |
[45] | Plimack ER , Dunbrack RL , Brennan TA , Andrake MD , Zhou Y , Serebriiskii IG , et al. Defects in DNA Repair Genes Predict Response to Neoadjuvant Cisplatin-based Chemotherapy in Muscle-invasive Bladder Cancer. Eur Urol (2015) ;68: (6):959–67. |
[46] | Groenendijk FH , de Jong J , Fransen van de Putte EE , Michaut M , Schlicker A , Peters D , et al. ERBB2 Mutations Characterize a Subgroup of Muscle-invasive Bladder Cancers with Excellent Response to Neoadjuvant Chemotherapy. Eur Urol (2015) ;69: (3):384–8. |
[47] | Van Allen EM , Mouw KW , Kim P , Iyer G , Wagle N , Al-Ahmadie H , et al. Somatic ERCC2 mutations correlate with cisplatin sensitivity in muscle-invasive urothelial carcinoma. Cancer Discov (2014) ;4: (10):1140–53. |
[48] | McConkey DJ , Choi W , Shen Y , Lee IL , Porten S , Matin SF , et al. A Prognostic Gene Expression Signature in the Molecular Classification of Chemotherapy-naive Urothelial Cancer is Predictive of Clinical Outcomes from Neoadjuvant Chemotherapy: A Phase 2 Trial of Dose-dense Methotrexate, Vinblastine, Doxorubicin, and Cisplatin with Bevacizumab in Urothelial Cancer. Eur Urol (2015) ;69: (5):855–62. |
[49] | Wagle N , Grabiner BC , Van Allen EM , Hodis E , Jacobus S , Supko JG , et al. Activating mTOR mutations in a patient with an extraordinary response on a phase I trial of everolimus and pazopanib. Cancer Discov (2014) ;4: (5):546–53. |
[50] | Iyer G , Hanrahan AJ , Milowsky MI , Al-Ahmadie H , Scott SN , Janakiraman M , et al. Genome sequencing identifies a basis for everolimus sensitivity. Science (2012) ;338: (6104):221–. |
[51] | Sequist LV CP , Varga A , et al. Phase I study of BGJ398, a selective pan-FGFR inhibitor in genetically preselected advanced solid tumors. American Association for Cancer Research Annual Meeting 2014; 5–9, (2014) ; San Diego Abstr CT326. |
[52] | Choudhury NJ , Campanile A , Antic T , Yap KL , Fitzpatrick CA , Wade JL 3rd , et al. Afatinib Activity in Platinum-Refractory Metastatic Urothelial Carcinoma in Patients With ERBB Alterations. J Clin Oncol (2016) ;34: (18):2165–71. |
[53] | Millner LM , Linder MW , Valdes R Jr . Circulating tumor cells: A review of present methods and the need to identify heterogeneous phenotypes. Ann Clin Lab Sci (2013) ;43: (3):295–304. |
[54] | Yun SJ , Kim WJ . Role of the epithelial-mesenchymal transition in bladder cancer: From prognosis to therapeutic target. Korean J Urol (2013) ;54: (10):645–50. |
[55] | Gallagher DJ , Milowsky MI , Ishill N , Trout A , Boyle MG , Riches J , et al. Detection of circulating tumor cells in patients with urothelial cancer. Ann Oncol (2009) ;20: (2):305–8. |
[56] | Flaig TW , Wilson S , van Bokhoven A , Varella-Garcia M , Wolfe P , Maroni P , et al. Detection of circulating tumor cells in metastatic and clinically localized urothelial carcinoma. Urology (2011) ;78: (4):863–7. |
[57] | Naoe M , Ogawa Y , Morita J , Omori K , Takeshita K , Shichijyo T , et al. Detection of circulating urothelial cancer cells in the blood using the CellSearch System. Cancer (2007) ;109: (7):1439–45. |
[58] | De Santis M . Circulating tumor cells (CTC) and HER-2 status on CTC and primary tumor in urothelial cancer (UC) patients refractory to platinum based chemotherapy. J Clin Oncol (2016) ;34: (suppl; abstr 4520). 2016. |
[59] | Guzzo TJ , McNeil BK , Bivalacqua TJ , Elliott DJ , Sokoll LJ , Schoenberg MP . The presence of circulating tumor cells does not predict extravesical disease in bladder cancer patients prior to radical cystectomy. Urol Oncol (2012) ;30: (1):44–8. |
[60] | Rink M , Chun FK , Dahlem R , Soave A , Minner S , Hansen J , et al. Prognostic role and HER2 expression of circulating tumor cells in peripheral blood of patients prior to radical cystectomy: A prospective study. Eur Urol (2012) ;61: (4):810–7. |
[61] | Gazzaniga P , de Berardinis E , Raimondi C , Gradilone A , Busetto GM , De Falco E , et al. Circulating tumor cells detection has independent prognostic impact in high-risk non-muscle invasive bladder cancer. Int J Cancer (2014) ;135: (8):1978–82. |
[62] | Gradilone A , Petracca A , Nicolazzo C , Gianni W , Cortesi E , Naso G , et al. Prognostic significance of survivin-expressing circulating tumour cells in T1G3 bladder cancer. BJU Int (2010) ;106: (5):710–5. |
[63] | Kinjo M , Okegawa T , Horie S , Nutahara K , Higashihara E . Detection of circulating MUC7-positive cells by reverse transcription-polymerase chain reaction in bladder cancer patients. Int J Urol (2004) ;11: (1):38–43. |
[64] | Li SM , Zhang ZT , Chan S , McLenan O , Dixon C , Taneja S , et al. Detection of circulating uroplakin-positive cells in patients with transitional cell carcinoma of the bladder. J Urol (1999) ;162: (3 Pt 1):931–5. |
[65] | Gazzaniga P , Gandini O , Giuliani L , Magnanti M , Gradilone A , Silvestri I , et al. Detection of epidermal growth factor receptor mRNA in peripheral blood: A new marker of circulating neoplastic cells in bladder cancer patients. Clin Cancer Res (2001) ;7: (3):577–83. |
[66] | Qi F , Liu Y , Zhao R , Zou X , Zhang L , Li J , et al. Quantitation of rare circulating tumor cells by folate receptor alpha ligand-targeted PCR in bladder transitional cell carcinoma and its potential diagnostic significance. Tumour Biol (2014) ;35: (7):7217–23. |
[67] | Fujii Y , Kageyama Y , Kawakami S , Kihara K , Oshima H . Detection of disseminated urothelial cancer cells in peripheral venous blood by a cytokeratin 20-specific nested reverse transcriptase-polymerase chain reaction. Jpn J Cancer Res (1999) ;90: (7):753–7. |
[68] | Lu JJ , Kakehi Y , Takahashi T , Wu XX , Yuasa T , Yoshiki T , et al. Detection of circulating cancer cells by reverse transcription-polymerase chain reaction for uroplakin II in peripheral blood of patients with urothelial cancer. Clin Cancer Res (2000) ;6: (8):3166–71. |
[69] | Alva A , Friedlander T , Clark M , Huebner T , Daignault S , Hussain M , et al. Circulating Tumor Cells as Potential Biomarkers in Bladder Cancer. J Urol (2015) ;194: (3):790–8. |
[70] | Premasekharan G , Gilbert E , Okimoto RA , Hamirani A , Lindquist KJ , Ngo VT , et al. An improved CTC isolation scheme for pairing with downstream genomics: Demonstrating clinical utility in metastatic prostate, lung and pancreatic cancer. Cancer Lett (2016) ;380: (1):144–52. |
[71] | Scher HI , Lu D , Schreiber NA , Louw J , Graf RP , Vargas HA , et al. Association of AR-V7 on Circulating Tumor Cells as a Treatment-Specific Biomarker With Outcomes and Survival in Castration-Resistant Prostate Cancer. JAMA Oncol (2016) . |
[72] | Antonarakis ES , Lu C , Wang H , Luber B , Nakazawa M , Roeser JC , et al. AR-V7 and resistance to enzalutamide and abiraterone in prostate cancer. N Engl J Med (2014) ;371: (11):1028–38. |
[73] | Necchi A , Fina E, Giannatempo P, et al. Quantification and molecular profiling of circulating tumor cells (CTCs) in urothelial cancer (UC) before and during systemic treatment: Implications across the clinical stages. J Clin Oncol (2014) ;32: (suppl4; abstr 319). 2014. |
[74] | Msaouel P , Koutsilieris M . Diagnostic value of circulating tumor cell detection in bladder and urothelial cancer: Systematic review and meta-analysis. BMC Cancer (2011) ;11: :336. |
[75] | Bettegowda C , Sausen M , Leary RJ , Kinde I , Wang Y , Agrawal N , et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci Transl Med (2014) ;6: (224):224ra24. |
[76] | Birkenkamp-Demtroder K , Nordentoft I , Christensen E , Hoyer S , Reinert T , Vang S , et al. Genomic Alterations in Liquid Biopsies from Patients with Bladder Cancer. Eur Urol (2016) ;70: (1):75–82. |
[77] | Nagy RJ , Agarwal N, Gupta S , et al. Circulating cell-free DNA profiling of patients with advanced urothelial carcinoma of the bladder. J Clin Oncol (2016) ;34: (suppl; abstr 4528). |
[78] | Henrique R , Costa VL , Jeronimo C . Methylation-based biomarkers for early detection of urological cancer. Crit Rev Oncog (2007) ;13: (4):265–82. |
[79] | Dominguez G , Carballido J , Silva J , Silva JM , Garcia JM , Menendez J , et al. p14ARF promoter hypermethylation in plasma DNA as an indicator of disease recurrence in bladder cancer patients. Clin Cancer Res (2002) ;8: (4):980–5. |
[80] | Ellinger J , El Kassem N , Heukamp LC , Matthews S , Cubukluoz F , Kahl P , et al. Hypermethylation of cell-free serum DNA indicates worse outcome in patients with bladder cancer. J Urol (2008) ;179: (1):346–52. |
[81] | Jablonowski Z , Reszka E , Gromadzinska J , Wasowicz W , Sosnowski M . Hypermethylation of p16 and DAPK promoter gene regions in patients with non-invasive urinary bladder cancer. Arch Med Sci (2011) ;7: (3):512–6. |
[82] | Valenzuela MT , Galisteo R , Zuluaga A , Villalobos M , Nunez MI , Oliver FJ , et al. Assessing the use of p16(INK4a) promoter gene methylation in serum for detection of bladder cancer. Eur Urol (2002) ;42: (6):622–8; discussion 8-30. |
[83] | Lin YL , Sun G , Liu XQ , Li WP , Ma JG . Clinical significance of CDH13 promoter methylation in serum samples from patients with bladder transitional cell carcinoma. J Int Med Res (2011) ;39: (1):179–86. |
[84] | Luo ZG , Li ZG , Gui SL , Chi BJ , Ma JG . Protocadherin-17 promoter methylation in serum-derived DNA is associated with poor prognosis of bladder cancer. J Int Med Res (2014) ;42: (1):35–41. |
[85] | Friedrich MG , Chandrasoma S , Siegmund KD , Weisenberger DJ , Cheng JC , Toma MI , et al. Prognostic relevance of methylation markers in patients with non-muscle invasive bladder carcinoma. Eur J Cancer (2005) ;41: (17):2769–78. |
[86] | Tam S , de Borja R , Tsao MS , McPherson JD . Robust global microRNA expression profiling using next-generation sequencing technologies. Lab Invest (2014) ;94: (3):350–8. |
[87] | Catto JW , Miah S , Owen HC , Bryant H , Myers K , Dudziec E , et al. Distinct microRNA alterations characterize high- and low-grade bladder cancer. Cancer Res (2009) ;69: (21):8472–81. |
[88] | Bellmunt J , Zhou CW , Mullane SA , Werner L , Taplin ME , Fay AP , et al. Association of tumour microRNA profiling with outcomes in patients with advanced urothelial carcinoma receiving first-line platinum-based chemotherapy. Br J Cancer (2016) ;115: (1):12–9. |
[89] | Zhang X , Zhang Y , Liu X , Fang A , Li P , Li Z , et al. MicroRNA-203 Is a Prognostic Indicator in Bladder Cancer and Enhances Chemosensitivity to Cisplatin via Apoptosis by Targeting Bcl-w and Survivin. PLoS One (2015) ;10: (11):e0143441. |
[90] | Guancial EA , Bellmunt J , Yeh S , Rosenberg JE , Berman DM . The evolving understanding of microRNA in bladder cancer. Urol Oncol (2014) ;32: (1):41 e31–40. |
[91] | Jiang X , Du L , Duan W , Wang R , Yan K , Wang L , et al. Serum microRNA expression signatures as novel noninvasive biomarkers for prediction and prognosis of muscle-invasive bladder cancer. Oncotarget (2016) ;7: (24):36733–36742. |
[92] | Jiang X , Du L , Wang L , Li J , Liu Y , Zheng G , et al. Serum microRNA expression signatures identified from genome-wide microRNA profiling serve as novel noninvasive biomarkers for diagnosis and recurrence of bladder cancer. Int J Cancer (2015) ;136: (4):854–62. |
[93] | Fang Z , Dai W , Wang X , Chen W , Shen C , Ye G , et al. Circulating miR- A promising biomarker for the detection and prognosis evaluation of bladder cancer. Tumour Biol (2016) ;37: (6):8075–82. |
[94] | Du M , Shi D , Yuan L , Li P , Chu H , Qin C , et al. Circulating miR-497 and miR-663b in plasma are potential novel biomarkers for bladder cancer. Sci Rep (2015) ;5: :10437. |
[95] | Matsumoto K , Shariat SF , Casella R , Wheeler TM , Slawin KM , Lerner SP . Preoperative plasma soluble E-cadherin predicts metastases to lymph nodes and prognosis in patients undergoing radical cystectomy. J Urol (2003) ;170: (6 Pt 1):2248–52. |
[96] | Svatek RS , Shah JB , Xing J , Chang D , Lin J , McConkey DJ , et al. A multiplexed, particle-based flow cytometric assay identified plasma matrix metalloproteinase-7 to be associated with cancer-related death among patients with bladder cancer. Cancer (2010) ;116: (19):4513–9. |
[97] | Shariat SF , Kim JH , Andrews B , Kattan MW , Wheeler TM , Kim IY , et al. Preoperative plasma levels of transforming growth factor beta(1) strongly predict clinical outcome in patients with bladder carcinoma. Cancer (2001) ;92: (12):2985–92. |
[98] | Shariat SF , Monoski MA , Andrews B , Wheeler TM , Lerner SP , Slawin KM . Association of plasma urokinase-type plasminogen activator and its receptor with clinical outcome in patients undergoing radical cystectomy for transitional cell carcinoma of the bladder. Urology (2003) ;61: (5):1053–8. |
[99] | Szarvas T , Laszlo V , Vom Dorp F , Reis H , Szendroi A , Romics I , et al. Serum endostatin levels correlate with enhanced extracellular matrix degradation and poor patients’ prognosis in bladder cancer. Int J Cancer (2012) ;130: (12):2922–9. |
[100] | Bellmunt J , Gonzalez-Larriba JL , Prior C , Maroto P , Carles J , Castellano D , et al. Phase II study of sunitinib as first-line treatment of urothelial cancer patients ineligible to receive cisplatin-based chemotherapy: Baseline interleukin-8 and tumor contrast enhancement as potential predictive factors of activity. Ann Oncol (2011) ;22: (12):2646–53. |
[101] | Necchi A , Mariani L , Zaffaroni N , Schwartz LH , Giannatempo P , Crippa F , et al. Pazopanib in advanced and platinum-resistant urothelial cancer: An open-label, single group, phase 2 trial. The Lancet Oncology (2012) ;13: (8):810–6. |
[102] | Schwamborn K , Krieg RC , Grosse J , Reulen N , Weiskirchen R , Knuechel R , et al. Serum proteomic profiling in patients with bladder cancer. Eur Urol (2009) ;56: (6):989–96. |
[103] | Cao M , Zhao L , Chen H , Xue W , Lin D . NMR-based metabolomic analysis of human bladder cancer. Anal Sci (2012) ;28: (5):451–6. |
[104] | Schiess R , Wollscheid B , Aebersold R . Targeted proteomic strategy for clinical biomarker discovery. Mol Oncol (2009) ;3: (1):33–44. |
[105] | Stadler WM , Lerner SP , Groshen S , Stein JP , Shi S-R , Raghavan D , et al. Phase III study of molecularly targeted adjuvant therapy in locally advanced urothelial cancer of the bladder based on p53 status. Journal of Clinical Oncology: Official Journal of the American Society of Clinical Oncology (2011) ;29: (25):3443–9. |
[106] | Lee JK , Havaleshko DM , Cho H , Weinstein JN , Kaldjian EP , Karpovich J , et al. A strategy for predicting the chemosensitivity of human cancers and its application to drug discovery. Proceedings of the National Academy of Sciences of the United States of America (2007) ;104: (32):13086–91. |
Figures and Tables
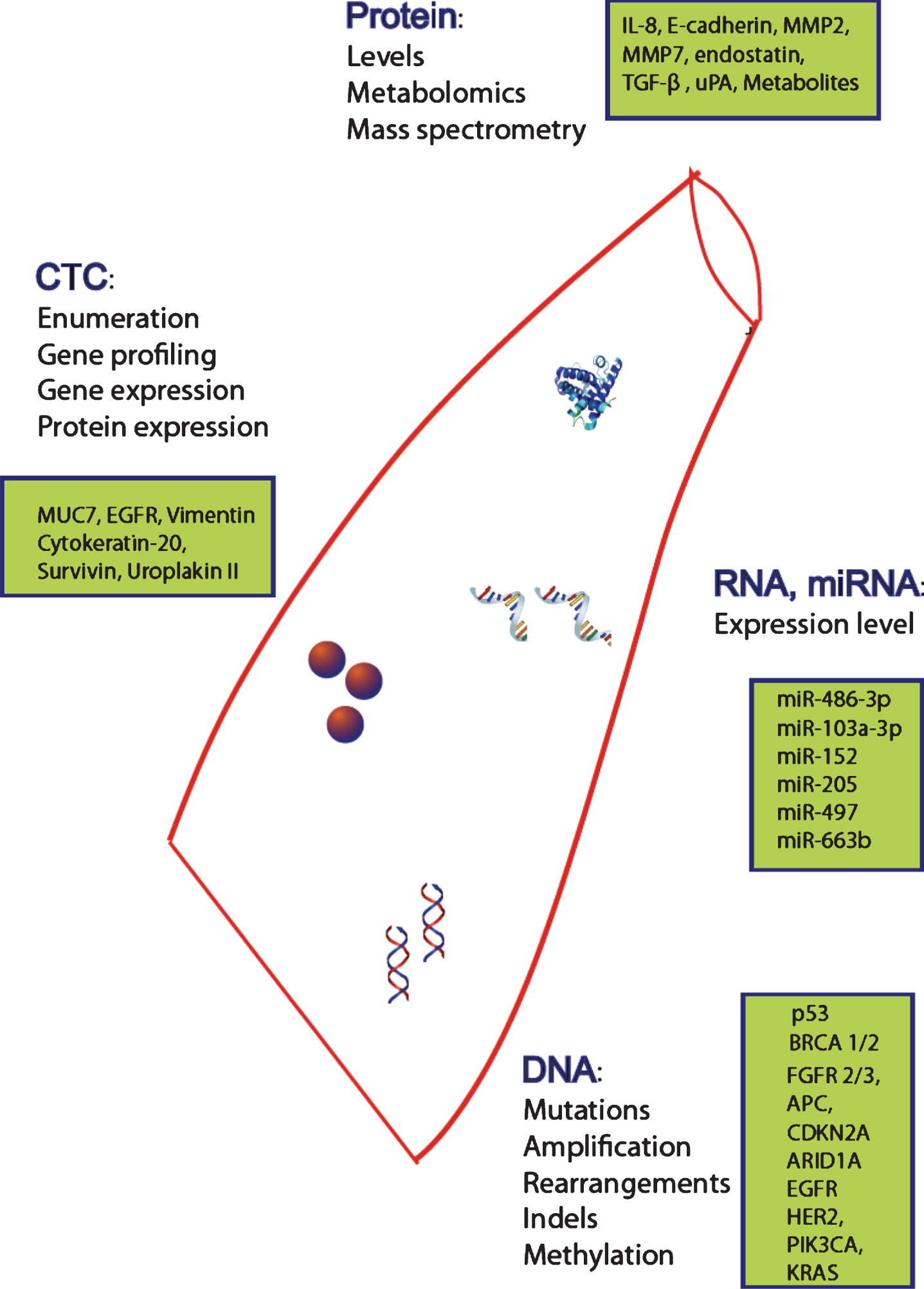
Fig.1
Potential circulating biomarkers. Potential circulating biomarkers for the management of bladder cancer including CTCs (circulating tumor cells) for enumeration and profiling, DNA, RNA, miRNA and protein (levels, metabolomics and spectrometry).