Design and first applications of a flexible Raman micro-spectroscopic system for biological imaging
Abstract
Typical commercial Raman micro-spectroscopic systems do not offer much flexibility to the end user, thus limiting potential research applications. We present a design of a simple, highly flexible and portable confocal Raman microscope with a detailed list of parts. The system can perform spectral acquisition in different modes: single-point spectroscopy, hyperspectral point mapping or hyperspectral line mapping. Moreover, the microscope can be easily converted between inverted and upright configurations, which can be beneficial for specific situations. Fiber coupling enables to connect various lasers for excitation and spectrometer/CCD combinations for signal detection. The performance of the instrument is demonstrated via Raman spectroscopy at 785 nm excitation wavelength, single point mapping of pancreatic cancer cells placed onto a quartz substrate and line mapping of polystyrene beads.
1.Introduction
Raman spectroscopy, a popular subtype of vibrational spectroscopy, evolved in last decades into a powerful technique for analysis and chemical characterization of specimens. A Raman spectrum results from inelastic light scattering on vibrating molecular bonds (see basic theory in Section 2), therefore the spectrum is specific for each particular substance and contains information about molecular composition of the specimen.
Ability to combine non-destructive chemical characterization with diffraction-limited lateral resolution makes Raman micro-spectroscopy a versatile research tool [45,50]. It is particularly well suited for biological research, because it allows to detect variations in the biomolecular composition and correlate them with the corresponding biological changes due to metabolism or pathology. Absence of lengthy sample preparation, such as staining or labeling, makes Raman spectroscopy a promising tool for rapid clinical diagnosis [7,23,26,29,44,52]. Not only biological tissues [27] and body fluids [18] can be examined by Raman spectroscopy, but even individual living cells [17,32,42]. This is a large research field with a host of promising applications, such as observation of cell metabolism, growth and aging, study of drug resistance or drug uptake [35], chemical mapping of cells [22,36], identification of cell in a mixed population, and many others. This could guide us to the understanding of changes within an individual cell that could lead to a disease development, because many diseases begin at the cellular level.
A range of commercial Raman microscopic systems exists. Usually, these are robust systems with built-in calibration routines and user-friendly software. However, they are often dedicated for a specific purpose and therefore have limited potential for modifications. Nonetheless, it is possible to assemble a confocal Raman microscope from off-the-shelf components. It can be further extended with special Raman detection schemes (e.g. line scanning-mode [21], modulation of the excitation wavelength [15], or dual-polarization [11]) and supplemented with additional detection techniques like fluorescence [25,31], phase contrast [25], dark-field [31] microscopy, and others [53]. Moreover, custom systems can be combined with advanced sample handling techniques, like microfluidics [2,3,14] or optical tweezers [1,24,25,57].
This contribution presents a detailed design of a Raman microscope including a scheme of optical layout and a list of components. We discuss important parameters of excitation laser, camera and spectrometer, as well as implementation of the Raman imaging. Mechanical stability and flexibility of the system, as well as spatial resolution of the microscope are considered as well. A set of hyperspectral confocal Raman images of two cancer cell lines has been acquired to show the performance of the instrument. Finally, we demonstrate fast Raman mapping of polystyrene beads in a line-scanning imaging mode.
2.Basic theory
Raman effect is the inelastic scattering of photons on vibrational modes of molecular bonds [33,48]. An incident photon with energy
The difference in energy between incoming and scattered photons
There are multiple variations of Raman effect, such as resonance Raman scattering, stimulated Raman scattering or coherent anti-Stokes Raman scattering [33,46], but this contribution deals only with spontaneous Stokes scattering.
3.Design of the Raman micro-spectroscopic system
A Raman detection system is supposed to collect Stokes-shifted scattered photons and guide them to the entrance slit of a spectrometer. As the inelastically scattered photons are emitted isotropically from the sample, an optical system is required that would collect them and refocus on the spectrometer input slit. An infinity-corrected microscope objective lens with high numerical aperture is particularly suitable for this purpose for several reasons. First, it provides high energy density for excitation in the focal spot, as well as gives a possibility to selectively probe small spots of a heterogeneous sample located at different positions. Second, it collects a large portion of the inelastically scattered photons emitted from the observation point due to its high acceptance angle. Finally, collected light leaves the objective lens as a collimated beam that can be easily focused on the spectrometer entrance slit with a tube lens.
In last decades the industry of optical filters underwent major leap forward, which revolutionized the way Raman systems are built and drastically improved their performance. Compact narrow-band optical filters superseded bulky monochromators used in past systems; dichroic filters, which offer higher transmission, replaced beam splitters/combiners. A notch dichroic beam splitter (NDBS) is a perfect option to couple the excitation laser beam into the optical microscope (see Fig. 1). Placed between the tube and objective lenses, it reflects the excitation light and directs it towards the objective lens. Additionally, it transmits the inelastically scattered light towards the spectrometer while suppressing the Rayleigh-scattered light.
Fig. 1.
Left – optical layout of the experimental setup. Two Raman detection paths are shown: free-space line-scanning via optical elements L4-6 and M3,4, as well as confocal detection via lens L4 and optical fiber MMF. See Table 1 for specifications of parts and abbreviations used throughout. Right – transmission spectra of optical filters. Spectral data provided by Semrock Inc.
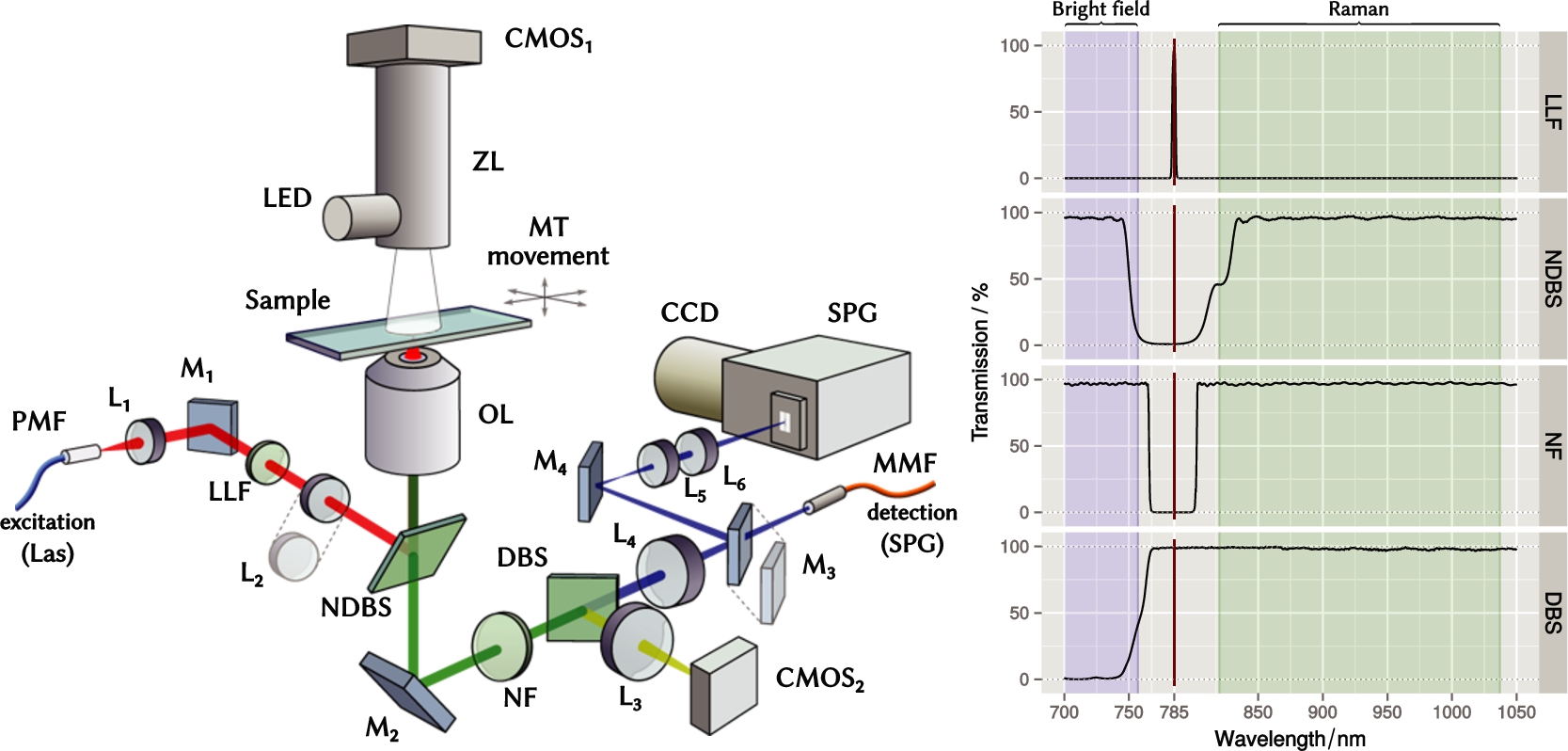
Table 1
Abbreviations of the equipment depicted in Fig. 1
| Abbr. | Description |
| CCD | Peltier-cooled back-illuminated deep depletion CCD camera PIXIS-400BR-eXcelon, |
| CMOS1,2 | Color CMOS camera DCC1645C, |
| DBS | Dichroic long-pass beam splitter FF757-Di01-25x36, 757 nm edge wavelength | Semrock Inc, USA |
| L1,3-6 | Plano-convex lenses with anti-reflection coating | Thorlabs Inc. |
| L2 | Non-axisymmetrical optical element, e.g. cylindrical lens or Powell lens |
| Las | Single mode laser XTRA, 785 nm wavelength, 450 mW optical power, line width below 10 MHz, | Toptica Photonics AG, Germany |
| LED | Cold white LED MCWHL5, 1000 mA current, 800 mW optical power | Thorlabs Inc. |
| LLF | Laser line filter, LL01-785 | Semrock Inc, USA |
| M1,2,4 | Silver-coated mirrors in tilt mounts |
| M3 | Removable silver-coated mirror in a tilt magnetic mount |
| MMF | Multimode optical fiber, 105 µm core diameter |
| MT | Motorized table, three motorized translation stages with 25 mm travel range. Lateral axes: two CONEX-MFACC | Newport Corp. Vertical axis: MTS25-Z8 | Thorlabs Inc. |
| NDBS | Notch dichroic beam splitter, NFD01-785 | Semrock Inc, USA |
| NF | Notch filter LC-785NF-25, 39 nm bandwidth | Laser Components GmbH, Germany |
| OL | Microscope objective lenses: Nikon 60× NA 1.0 water-immersion or Nikon 100× NA 1.4 oil-immersion, in the revolver objective turret (OT1 | Thorlabs Inc.) |
| PMF | Polarization-maintaining optical fiber with FC/APC ferrules | Thorlabs Inc. |
| SPG | Compact imaging spectrograph IsoPlane 160, |
| ZL | High magnification 12× zoom lens | Navitar Inc., USA |
Many optical microscopes can be upgraded to Raman microscopes by adding several additional components [10]. It is important to note that it is preferable to use an infinity-corrected optical system that features a parallel optical path between the objective lens and the tube lens. All additional components, such as filters, polarizers, prisms, etc., can be placed into this parallel path without introducing additional spherical aberrations [30]. Modern research-grade biomedical and industrial microscopes typically fall into the category of infinity-corrected optical systems.
3.1.Spatial resolution
The spatial resolution of a microscope is determined primarily by the objective lens that focuses light into a diffraction-limited spot. The spatial resolution, especially along the optical axis, can be further enhanced by using a confocal detection approach, i.e. the collected light is focused and transmitted through a pinhole located in front of the detector. The pinhole is optically conjugated with the interrogation volume, from which the Raman spectrum is acquired, such as it accepts only the light originating from the location of the laser spot and rejects the rest [49,55]. It is convenient to guide the signal to the spectrometer using a multimode optical fiber. As the fiber core acts the same way as a pinhole placed in the back focal plane of the microscope, a smaller fiber diameter improves the confocality.
Confocal Raman system allows to “look inside” a semi-transparent sample and probe its chemical composition at a particular depth. The biggest advantage of the confocal detection scheme is the suppression of the unwanted signal originating from the background (i.e. specimen locations adjacent to the interrogation volume). However, since the pinhole rejects the out-of-plane light, less signal is collected by the detector. Therefore, it is necessary to find a reasonable trade-off between the resolution and the sensitivity of the instrument. The confocality of the system is a function of several factors, such as objective lens, tube lens, and the diameter of the detection fiber core. A rigorous assessment of the confocal Raman microscopy can be found elsewhere [19,30,49].
3.2.Implementation of hyperspectral imaging
An important design question is the implementation of scanning, which is required for acquisition of hyperspectral Raman images. One of the options considered by us was laser scanning (LS), i.e. steering of the laser beam by a pair of mirror galvanometers [20,34].
Albeit LS system features fast operation, and successful implementations of Raman LS microscope have been reported [47,54], such a system has several inherent drawbacks. The key problem is vignetting, i.e. the deterioration of the Raman collection efficiency when the excitation spot goes off-axis. Other issues are limited field of view, as well as reduced light transmission and complicated alignment due to a high number of optical components.
Other option, which is free from the aforementioned issues, is the movement of the specimen across the fixed laser focus. In our system we use three programmable motorized linear stages, attached orthogonally to each other (see Table 1 for details). They feature 25 mm travel range with resolution down to 100 nm, thus enabling precise and reproducible sample positioning under the microscope objective.
3.3.Considerations for the excitation laser
The excitation laser is an essential component of any Raman spectroscopy system. The main relevant laser parameters are power, wavelength, line width and stability.
The Raman spectrum represents the energy difference between excitation photons and modes of molecular vibrations, as such any uncertainty or deviation of the laser wavelength directly affects the wavenumber position and, consequently, the spectral resolution. Therefore, it is important to ensure that the laser has a narrow emission line, and its wavelength is well-defined. Otherwise, the wavelength uncertainty would lead to an additional broadening of the detected spectrum.
Another important factor is the wavelength stability over time. Jumping laser wavelength would render the wavenumber calibration of an instrument invalid and make evaluation of the spectral data cumbersome.
Typically a so-called single frequency laser is used as an excitation source, which has a single longitudinal mode and thus features a very small line width, low phase noise and low intensity variations. Such lasers are very sensitive to optical feedback, so they should be carefully protected from any back-reflections, using for example a Faraday optical isolator and fiber optic connectors with pre-angled ferrules (FC/APC type).
The choice of laser power and wavelength, however, is not straightforward and is dictated by several factors [48]. The intensity of the Raman scattering is linearly proportional to the laser power, but it depends on the fourth power of the excitation light frequency, hence shorter wavelength provides higher Raman signal. However, with a short excitation wavelength a lot of undesired effects can occur. Photons of ultraviolet and visible light are capable of causing electronic excitation within molecules, which may lead to changes in chemical bonds, and consequently, to photo-degradation of a biological specimen. For example, even 5 mW power of green 514 nm laser beam suffice to cause a noticeable damage to a lymphocyte cell within several minutes; the process is accompanied by change of the Raman spectrum [40]. In contrast, it has been shown that bacterial cells can survive a prolonged exposure to 100 mW NIR excitation (790–1064 nm), which is long enough for the acquisition of a Raman spectrum [39]. An additional issue associated with the excitation laser emitting in the visible range is the autofluorescence of most biological samples. It is typically excited by wavelengths below 520 nm, and the emitted light has wavelength in the 450–600 nm range [4].
Too long excitation wavelength would be problematic as well. In addition to the decrease of scattering intensity with the forth power of the excitation frequency according to the Rayleigh law, one has to consider a rapid drop in the quantum efficiency of typical CCD detectors at wavelengths above 900–1000 nm. The sensor should have enough sensitivity to detect Stokes photons corresponding to the spectral features of interest. Since we are dealing with biological materials that typically exhibit complex spectral features used for identification in the so-called “fingerprint region” (approx. 700–1650 cm−1), it is important to ensure high detector sensitivity in the corresponding spectral range. With 785 nm excitation used in our case, the fingerprint region stretches up to 900 nm.
Another common region of interest is 2800–3000 cm−1. Energies of vibrational modes of C-Hx molecular bonds are located in this spectral region, which corresponds to 1006–1027 nm with 785 nm excitation. Typical CCD cameras have insufficient sensitivity above 1000 nm, so a special NIR-coated CCD sensor with enhanced quantum efficiency and suppressed etaloning is necessary.
Taking all aforementioned factors into account, we can conclude that the optimal excitation wavelength for Raman spectroscopy of biological materials would typically lie in red or NIR region.
3.4.Optical layout
The detailed optical layout of the Raman system is shown in Fig. 1 (see Table 1 for specifications of parts and abbreviations used throughout). The single mode excitation laser Las (Xtra, Toptica, Germany) provides up to 450 mW of optical power at wavelength of 785 nm with line width below 10 MHz. The laser is coupled to the microscopy setup via a polarization-maintaining fiber PMF. The PMF ensures that the polarization of the laser beam cannot fluctuate during the experiment. The laser beam is collimated using a lens L1 (
The laser line filter LLF, also called a plasma filter or laser clean-up filter, is used to remove any background and emission lines other than the main exciting line of the laser. It is important to note, that the Raman effect also happens inside of the PMF that delivers excitation light to the microscope, because the high-intensity laser light scatters on the vibrational modes of the quartz (the material of the fiber core). In this case the LLF filter becomes absolutely necessary.
The notch dichroic beam splitter NDBS couples the laser beam into the optical microscope and directs it toward the objective lens OL, which focuses the beam on the sample. The sample sits on a three-axes motorized table MT.
3.4.1.Detection of the Stokes signal
The scattered light, as well as the white illumination from a LED source, are collected by the same objective lens OL and sent through three optical filters: NDBS, NF, and DBS (see Fig. 1). The notch filter NF rejects the remaining Rayleigh-scattered laser light, while the dichroic beam splitter DBS separates the visible light, which is used for bright field imaging, from the Raman signal, which lies in the near-infrared range. The tube lens L4 focuses the parallel beam of Raman signal. Depending on the mode of operation, the Raman signal is delivered to the SPG either via MMF (confocal Raman) or free-space via mirrors M3,4 and lenses L5,6.
For confocal Raman micro-spectroscopy, the MMF is used to couple signal to the spectrometer. The fiber ferrule is placed into the image plane of the microscope – this way the fiber core acts as a pinhole, and the sample is scanned point-by-point to acquire a hyperspectral Raman image.
In an alternative configuration, the Raman signal is coupled to the spectrograph via free-space optical elements. In this case the signal collection is not confocal, so more light is collected, which enables faster spectrum acquisition at cost of decreased spatial resolution. The free-space coupling configuration allows to acquire Raman spectrum by fast laser line scanning. To do this, a non-axisymmetrical optical element L2 is introduced into the laser beam path to create a line-shaped focus. This focus is optically conjugated with the spectrometer slit, so that each line of the CCD camera sensor acquires an independent spectrum from an individual sample spot. In this way fast hyperspectral Raman imaging is performed by scanning the sample in one lateral direction only [43]. With the CCD camera used in our system it is possible to record up to 400 spectra simultaneously. The demonstration of Raman imaging with a laser line is given in Section 5. Lenses L5, L6 can be placed before the spectrometer to additionally magnify the image on the CCD sensor.
Hyperspectral laser-line imaging makes high demands on the imaging capabilities of the spectrograph (SPG). We used Princeton Instruments IsoPlane 160, which features aspherical optics that greatly reduces astigmatism and coma aberrations at all wavelengths across the entire focal plane. This makes the spectrograph particularly suitable for multichannel spectroscopy, in our case the Raman line-imaging, due to the reduced cross-talk. Additionally, sharp focusing yields higher signal-to-noise ratios for single spectral features.
The CCD camera installed in the system features low noise level and high quantum efficiency exceeding 90% across the 785–900 nm range, which is beneficial for fast data acquisition. Another important feature is the reduced etaloning (fringing), which enhances quality of the spectra acquired in the NIR region.
3.5.How to design the mechanical part
We used a computer-aided design (CAD) software (Autodesk Inventor) to create a computer model of the Raman system. We chose this approach for a number of reasons:
(1) Many suppliers of optomechanical components provide three-dimensional computer models of their products in the STEP format. These models can be downloaded and virtually assembled in the CAD software.
(2) The generated model clearly shows how the end system should look like. It is also possible to check position of every optical element and to measure any distance between arbitrary elements.
(3) Fast generation of documentation, including drawings, illustrations and bill of materials.
We used the so-called cage system to mount optical components, and we consider it to be the optimal choice for this task. Optical elements are firmly attached to aluminum plates that are connected together by rigid steel rods, which define a common optical axis, thus simplifying the alignment. The cage system enables modular design of the laboratory setup: different modules (e.g. detection module, laser coupling module, etc.) can be assembled and aligned independently of each other, and afterwards combined together. This also allows to quickly change the functionality of the instrument. Since the modules are small, rigid and stable, the setup can be flipped over in order to switch between upright and inverted configurations.
Usually upright and inverted microscopes are dedicated for different purposes and applications. Upright microscope, where the objective lens is located above the specimen, is well-suited for Raman spectroscopy of detached cells. The cells sink down to the substrate surface (e.g. CaF2 or quartz), then it is possible to locate and observe them with a physiological water immersion objective lens.
Inverted microscope configuration allows to examine live specimens in a deep covered container, such as a flask [9]. It is also advantageous for micro-manipulation applications, such as Raman-based flow cytometry [13,14,16,37], because it provides a lot of space to install additional tools. In our case, it enables good direction of the tubings towards a microfluidic chip, thus preventing sedimentation of the cells on tube walls. Additionally, we installed a 12× zoom lens ZL with CMOS1 camera above the specimen. The lens features large adjustable field of view and thus simplifies the navigation.
The mechanical stability is a particularly important parameter of any microscopy system. Since the excitation laser focus and the pinhole in front of the detector are conjugate points, even a small mechanical movement of the optical elements in the laser beam path can lead to the significant deterioration of the Raman system performance. An additional issue is the vibrational resistance: already a minor shock to the motorized table can lead to an unpredictable movement of the specimen under the objective. For this reason, the microscope modules, which are already built using the rigid case system, are attached to a 40 mm thick steel optical breadboard. The breadboard itself is suspended on top of four cushioning isolators filled with compressed air. The air pressure in the isolators is monitored and controlled by an active self-leveling electronics (PWA090, Thorlabs). This configuration mechanically decouples the measurement system from the environment and effectively cushions any external impacts.
4.Application to the study of cancer cells
We demonstrated performance of the custom-built Raman system by collecting Raman maps of individual fixed cells of two different cell lines: MIA PaCa-2, pancreatic cancer, and Jurkat, T lymphocytes. In total we acquired Raman maps of 10 individual cells of each cell line; each Raman map has size of
The cells were placed onto a 170 µm-thick quartz cover slip, and Raman signal has been collected in the inverted configuration. We used the 100× NA 1.4 oil-immersion objective lens both for Raman excitation and for Raman signal collection in 180° back reflection geometry. All scans were carried out with 100 mW power of the excitation laser (measured in the sample plane), exposure time of 1.0 second,
In total, 8000 Raman spectra have been collected in two batches. The spectra underwent preprocessing, which is a crucial step prior the data interpretation [8]. We used hyperSpec [6] package for programming language R [41] to perform all data manipulations, including preprocessing, analysis and visualization.
First of all, we calibrated the wavenumber axis in each of the batches using the standard calibration routine [51]. A spectrum of a Ne–Ar gas discharge lamp has been acquired, and observed peaks (about 45) have been mapped onto the atomic emission lines of Ne and Ar, provided by NIST [28]. The relationship between the CCD pixel number n and light wavelength
In order to reduce the fixed-pattern noise and to remove the constant ADC offset, we subtracted a dark frame from each spectrum. The dark frame has been collected by reading out the CCD camera while the shutter was closed. At the next step cosmic spikes have been detected and removed. After that, data from both batches have been down-sampled using LOESS local non-parametric regression onto a common wavenumber axis that has 240 points (initially 1340) and ranges from 550–3100 cm−1 . As a side effect, this lead to about two-fold increase in the signal-to-noise ratio, as well as a considerable reduction of the data size.
Each individual Raman map has been separated into cell and background regions using k-means clustering. An average spectrum of the background cluster has been calculated and subtracted from the Raman map. The remainder baseline has been estimated as a convex hull of the bent spectrum. Finally, all spectra have been intensity-corrected using a standardized incandescent lamp as a reference light source with known emission spectrum.
After the preprocessing, only the cell signatures, random noise and some tiny baseline fluctuations were left in the cell cluster. Figure 2 compares raw dataset with the preprocessed one.
Fig. 2.
Statistical overview of the collected spectral data. Lines represent, from bottom to top, the 5th, 16th, 50th (median), 84th, and 95th percentiles of the spectral intensity, respectively. The original raw dataset (8000 spectra, top) underwent smoothing interpolation, background removal, baseline subtraction and correction for the instrument response function. Then it was split into two subsets, containing spectra of MIA PaCa-2 and Jurkat cell lines, respectively. Numbers 1–14, described in Table 2, show commonly used bands for the assessment of biochemical composition. Note on scales: (1) Upper plot has a different scale. (2) The height of the peak located at 2700–3100 cm−1 in the preprocessed spectral data is reduced four times for the sake of visibility.
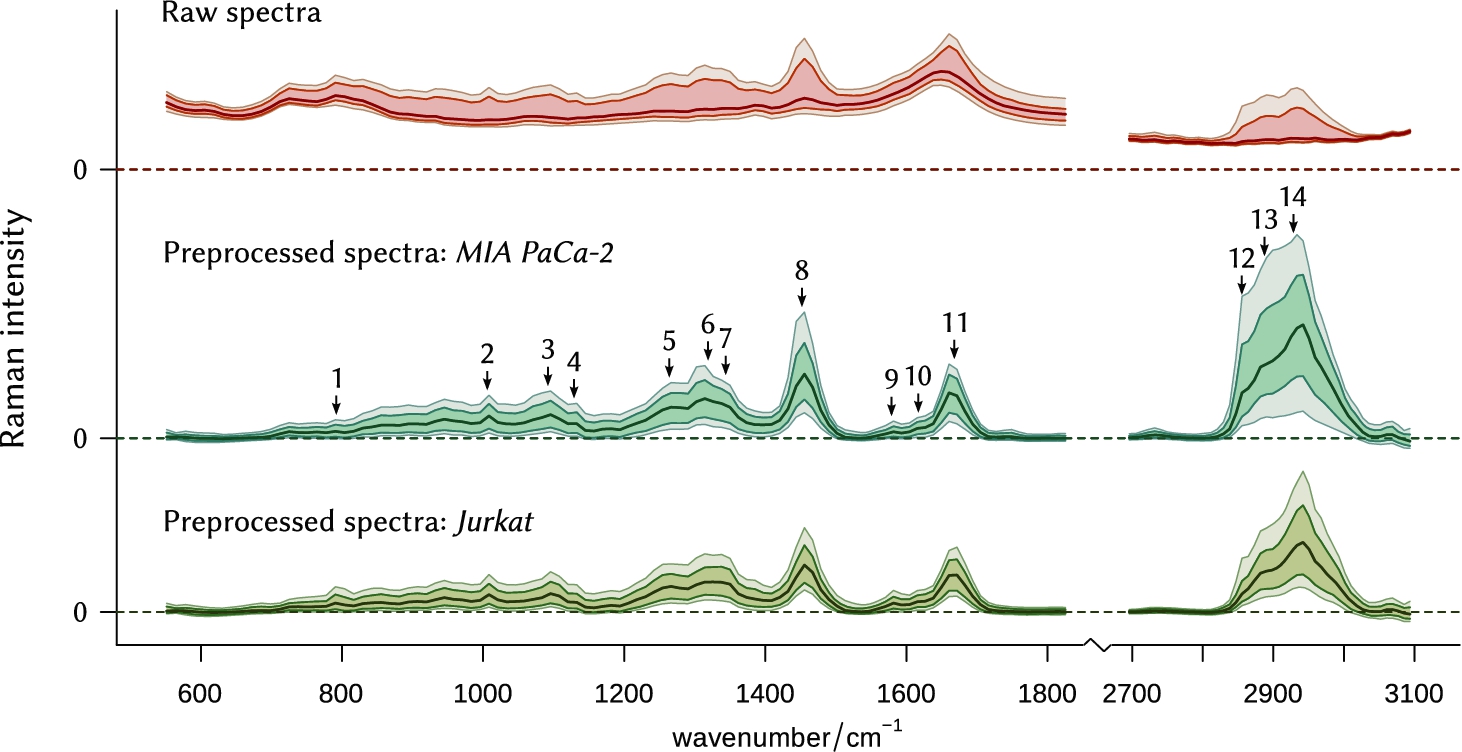
Table 2
| # | Wavenumber/cm−1 | Description* |
| 1 | 785–788 | na: C, T, U, |
| 2 | 1004 | p: Phe |
| 3 | 1090 | na: |
| 4 | 1127 | p: C–N str, lip: C–C str, |
| 5 | 1262 | na: G, p: amide III |
| 6 | 1319 | na: G, p: CH def, lip: CH2, CH3 twist |
| 7 | 1341 | na: A, G, p: CH def, |
| 8 | 1451 | p, lip: –CH2– def |
| 9 | 1585 | na: A, G |
| 10 | 1619 | p: Tyr, Trp |
| 11 | 1662 | na: T, p: amide I β-sheet, lip: fatty acids |
| 12 | 2855 | p, lip: –CH2– sym str |
| 13 | 2889 | p, lip: Fermi res. overtone of #8 |
| 14 | 2930 | p, lip: –CH2– asym str |
* Abbreviations: str: stretching; def: deformation vibration; sym: symmetric; asym: asymmetric; bk: backbone chain; na: nucleic acid; U, C, T, A, G: ring breathing modes of the DNA/RNA bases; p: protein; Tyr: tyrosine; Trp: tryptophan; Phe: phenylalanine; lip: lipid.
The N-FINDR algorithm [56] has been applied to the whole preprocessed dataset to extract three end-members, or the most different spectra. One of them is the blank background spectrum collected from points around the cell. Two other components, shown in Fig. 3(A), resemble spectra of lipid and DNA/protein mixture, respectively. We calculated concentration maps of these end-members for both cell lines (see Fig. 3(B)).
Fig. 3.
(A) – N-FINDR end-members that represent spectra of lipid and protein/DNA mixture. (B) – concentration maps of the end-members in cells of Jurkat and MIA PaCa-2 cell lines.
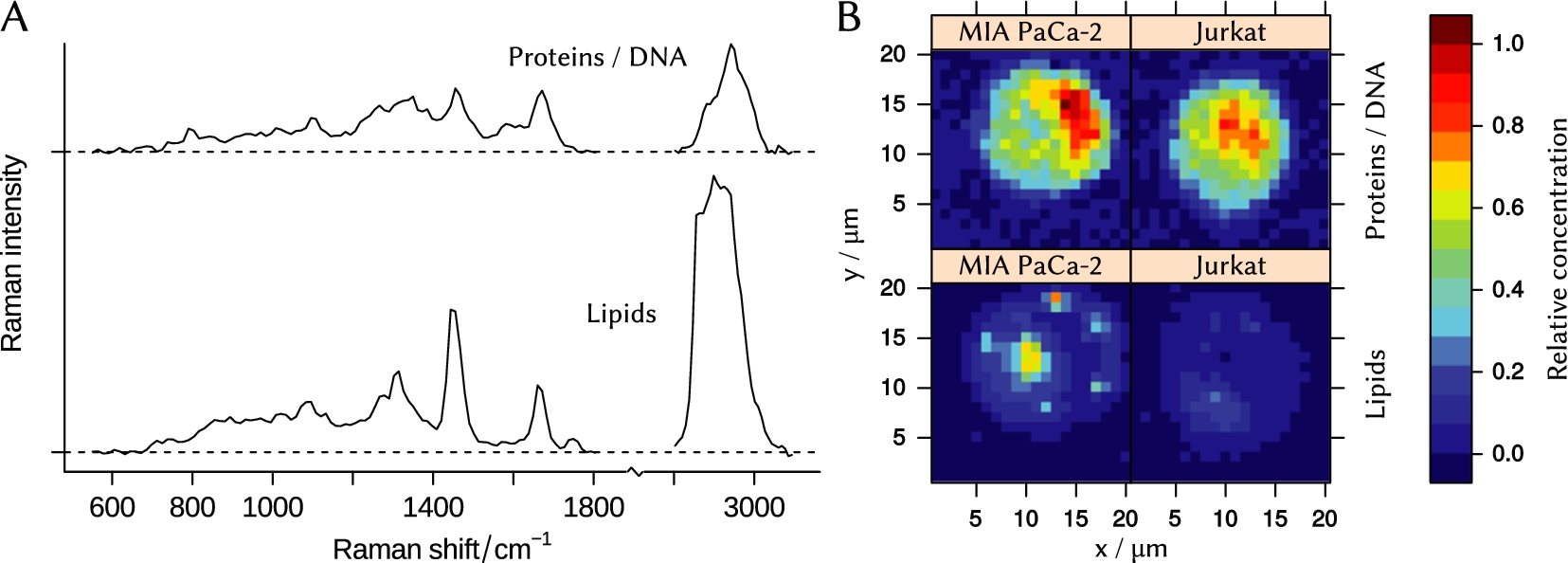
Results clearly show, that a typical MIA PaCa-2 cell contains more lipids compared to a typical Jurkat cell. The concentration of proteins differs as well. Obtained data can be used for purposes of cell identification [7,38] or planning of further biospectroscopic experiments [5].
5.Demonstration of line scanning
The performance of the Raman system in the hyperspectral laser line imaging mode has been tested on 5 µm polystyrene beads located on a CaF2 slide (see Fig. 4).
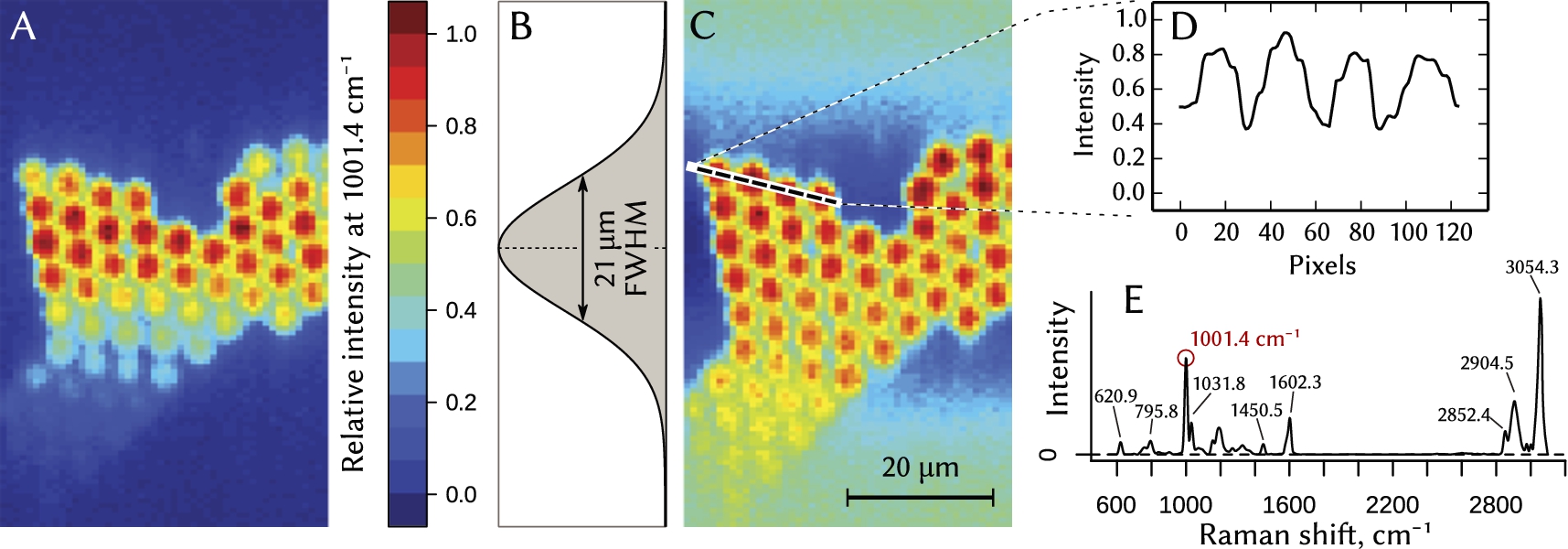
We placed a cylindrical lens L2 (see Fig. 1) into the laser beam to perform a one-dimensional beam shaping, i.e. to create an anamorphic beam that has different divergences in two mutually orthogonal planes. The beam diameter has been matched with the entrance aperture of the 60× NA 1.0 microscope objective lens OL, producing a thin linear-shaped focus with a diffraction-limited thickness of about 0.4 µm. The intensity profile of the line was close to Gaussian with 21 µm full width at half-maximum (Fig. 4B).
Raman signal has been collected from the area illuminated by the line-shaped focused beam in such a way, that each point along the linear focus corresponded to a specific location on the entrance slit of the spectrograph SPG. The line-shaped focus has been scanned across the sample in a single lateral direction in 1 µm steps. After each exposure of 0.5 seconds the CCD sensor has been read out, and the frame has been saved, followed by a movement to the next position on the specimen.
Use of the 60× objective lens with an
Fig. 4.
Hyperspectral Raman imaging of polystyrene beads with 5 µm diameter obtained in the line-scanning mode. False-color maps (A) and (C) show relative amplitude of the 1001.4 cm−1 peak in the Raman spectrum of the polystyrene (plot (E)). Raw data from CCD (image (A)) underwent intensity normalization (image (C)) using Gaussian intensity profile of the illumination spot (plot (B)) as a normalization factor. (D) – intensity profile along the dashed line in the normalized image (C).
6.Conclusion
We demonstrated the construction of a very flexible Raman micro-spectroscopic system for specific needs, in our case for the Raman-assisted cell identification. The described system features several acquisition modalities, in particular bright-field imaging, acquisition of Raman spectra from a single point, Raman mapping and hyperspectral line imaging.
Since the major part of the system consists of separate general purpose optomechanical components, it can be rebuilt, modified or extended. For example, a combination with fluorescence imaging requires an additional light source, a set of optical filters and a suitable camera for detection. Moreover, the whole microscopy attachment can be flipped, so one can switch between upright and inverted configurations.
We demonstrated the performance of the instrument by acquisition of hyperspectral Raman images in two different modes: point-by-point mapping and laser line scanning.
Acknowledgements
This work was supported by the EU within the FP7 collaborative project CanDo (610472), as well as by the Federal Ministry of Education and Research (BMBF) within the project RamanCTC (13N12685). Sonja Aškrabić was supported by a short time scientific mission within the COST action Raman4clinics (BM1401).
References
1 | [[1]] K. Ajito, C. Han and K. Torimitsu, Detection of glutamate in optically trapped single nerve terminals by Raman spectroscopy, Analytical Chemistry 76: (9) ((2004) ), 2506–2510. |
2 | [[2]] P.C. Ashok, G.P. Singh, H.A. Rendall, T.F. Krauss and K. Dholakia, Waveguide confined Raman spectroscopy for microfluidic interrogation, Lab. on a Chip 11: (7) ((2011) ), 1262. |
3 | [[3]] P.C. Ashok, G.P. Singh, K.M. Tan and K. Dholakia, Fiber probe based microfluidic Raman spectroscopy, Opt. Express 18: (8) ((2010) ), 7642–7649. |
4 | [[4]] J.E. Aubin, Autofluorescence of viable cultured mammalian cells, Journal of Histochemistry & Cytochemistry 27: (1) ((1979) ), 36–43. |
5 | [[5]] C. Beleites, U. Neugebauer, T. Bocklitz, C. Krafft and J. Popp, Sample size planning for classification models, Anal. Chim. Acta 760: ((2013) ), 25–33. |
6 | [[6]] C. Beleites and V. Sergo, hyperSpec: A package to handle hyperspectral data sets in R, available at: http://hyperspec.r-forge.r-project.org [January 13 2016], 2015. |
7 | [[7]] N. Bergner, T. Bocklitz, B.F.M. Romeike, R. Reichart, R. Kalff, C. Krafft and J. Popp, Identification of primary tumors of brain metastases by Raman imaging and support vector machines, in: Chemom. Intell. Lab. Syst., (2012) . |
8 | [[8]] T. Bocklitz, A. Walter, K. Hartmann, P. Rösch and J. Popp, How to pre-process Raman spectra for reliable and stable models?, Anal. Chim. Acta 704: (1,2) ((2011) ), 47–56. |
9 | [[9]] A.B. Zoladek, R.K. Johal, S. Garcia-Nieto, F. Pascut, K.M. Shakesheff, A.M. Ghaemmaghami and I. Notingher, Label-free molecular imaging of immunological synapses between dendritic and T cells by Raman micro-spectroscopy, The Analyst 135: (12) ((2010) ), 3205. |
10 | [[10]] P.J. Caspers, G.W. Lucassen and G.J. Puppels, Combined in vivo confocal Raman spectroscopy and confocal microscopy of human skin, Biophysical Journal 85: (1) ((2003) ), 572–580. |
11 | [[11]] L. Chiu, A.F. Palonpon, N.I. Smith, S. Kawata, M. Sodeoka and K. Fujita, Dual-polarization Raman spectral imaging to extract overlapping molecular fingerprints of living cells, Journal of Biophotonics 8: (7) ((2015) ), 546–554. |
12 | [[12]] J. De Gelder, K. De Gussem, P. Vandenabeele and L. Moens, Reference database of Raman spectra of biological molecules, J. Raman Spectrosc. 38: (9) ((2007) ), 1133–1147. |
13 | [[13]] S. Dochow, M. Becker, R. Spittel, C. Beleites, S. Stanca, I. Latka, K. Schuster, J. Kobelke, S. Unger, T. Henkel, G. Mayer, J. Albert, M. Rothhardt, C. Krafft and J. Popp, Raman-on-chip device and detection fibres with fibre Bragg grating for analysis of solutions and particles, Lab. on a Chip 13: (6) ((2013) ), 1109. |
14 | [[14]] S. Dochow, C. Beleites, T. Henkel, G. Mayer, J. Albert, J. Clement, C. Krafft and J. Popp, Quartz microfluidic chip for tumour cell identification by Raman spectroscopy in combination with optical traps, Analytical and Bioanalytical Chemistry 405: (8) ((2013) ), 2743–2746. |
15 | [[15]] S. Dochow, N. Bergner, C. Krafft, J. Clement, M. Mazilu, B.B. Praveen, P.C. Ashok, R. Marchington, K. Dholakia and J. Popp, Classification of Raman spectra of single cells with autofluorescence suppression by wavelength modulated excitation, Analytical Methods 5: (18) ((2013) ), 4608. |
16 | [[16]] S. Dochow, C. Krafft, U. Neugebauer, T. Bocklitz, T. Henkel, G. Mayer, J. Albert and J. Popp, Tumour cell identification by means of Raman spectroscopy in combination with optical traps and microfluidic environments, Lab. on a Chip 11: (8) ((2011) ), 1484. |
17 | [[17]] K. Eberhardt, C. Stiebing, C. Matthäus, M. Schmitt and J. Popp, Advantages and limitations of Raman spectroscopy for molecular diagnostics: An update, Expert Review of Molecular Diagnostics 15: (6) ((2015) ), 773–787. |
18 | [[18]] A.M.K. Enejder, T.G. Scecina, J. Oh, M. Hunter, W. Shih, S. Sasic, G.L. Horowitz and M.S. Feld, Raman spectroscopy for noninvasive glucose measurements, J. Biomed. Opt. 10: (3) ((2005) ), 031114-1–031114-9. |
19 | [[19]] N.J. Everall, Modeling and measuring the effect of refraction on the depth resolution of confocal Raman microscopy, Appl. Spectrosc. 54: (6) ((2000) ), 773–782. |
20 | [[20]] E. Fällman and O. Axner, Design for fully steerable dual-trap optical tweezers, Applied Optics 36: (10) ((1997) ), 2107–2113. |
21 | [[21]] L. Gao and R.T. Smith, Optical hyperspectral imaging in microscopy and spectroscopy – a review of data acquisition, Journal of Biophotonics 8: (6) ((2015) ), 441–456. |
22 | [[22]] M. Hedegaard, C. Matthäus, S. Hassing, C. Krafft, M. Diem and J. Popp, Spectral unmixing and clustering algorithms for assessment of single cells by Raman microscopic imaging, Theoretical Chemistry Accounts 130: (4–6) ((2011) ), 1249–1260. |
23 | [[23]] Z. Huang, G. Chen, X. Chen, J. Wang, J. Chen, P. Lu and R. Chen, Rapid and label-free identification of normal spermatozoa based on image analysis and micro-Raman spectroscopy, Journal of Biophotonics 7: (9) ((2014) ), 671–675. |
24 | [[24]] P.R.T. Jess, V. Garcés-Chávez, D. Smith, M. Mazilu, L. Paterson, A. Riches, C.S. Herrington, W. Sibbett and K. Dholakia, Dual beam fibre trap for Raman micro-spectroscopy of single cells, Opt. Express 14: (12) ((2006) ), 5779–5791. |
25 | [[25]] L. Kong, P. Zhang, G. Wang, J. Yu, P. Setlow and Y. Li, Characterization of bacterial spore germination using phase-contrast and fluorescence microscopy, Raman spectroscopy and optical tweezers, Nature Protocols 6: (5) ((2011) ), 625–639. |
26 | [[26]] C. Krafft and J. Popp, Raman4clinics: The prospects of Raman-based methods for clinical application, Analytical and Bioanalytical Chemistry 407: (27) ((2015) ), 8263–8264. |
27 | [[27]] C. Krafft, G. Steiner, C. Beleites and R. Salzer, Disease recognition by infrared and Raman spectroscopy, Journal of Biophotonics 2: (1,2) ((2009) ), 13–28. |
28 | [[28]] A. Kramida, Yu. Ralchenko, J. Reader and NIST ASD Team, NIST atomic spectra database (version 5.3), available at: http://physics.nist.gov/asd [January 13 2016], 2015. |
29 | [[29]] E.E. Lawson, B.W. Barry, A.C. Williams and H.G.M. Edwards, Biomedical applications of Raman spectroscopy, J. Raman Spectrosc. 28: (2,3) ((1997) ), 111–117. |
30 | [[30]] I.R. Lewis and H. Edwards, Handbook of Raman Spectroscopy: From the Research Laboratory to the Process Line, CRC Press, (2001) , p. 00379. |
31 | [[31]] H. Li, H. Wang, D. Huang, L. Liang, Y. Gu, C. Liang, S. Xu and W. Xu, Note: Raman microspectroscopy integrated with fluorescence and dark field imaging, Review of Scientific Instruments 85: (5) ((2014) ), 056109. |
32 | [[32]] Q. Li, E. Suasnavas, L. Xiao, S. Heywood, X. Qi et al., Label-free and non-invasive monitoring of porcine trophoblast derived cells: Differentiation in serum and serum-free media, Journal of Biophotonics 8: (8) ((2015) ), 638–645. |
33 | [[33]] D.A. Long, The Raman Effect: A Unified Treatment of the Theory of Raman Scattering by Molecules, Wiley, Chichester; New York, (2002) . |
34 | [[34]] G.F. Marshall, Handbook of Optical and Laser Scanning, Marcel Dekker, New York, (2004) . |
35 | [[35]] C. Matthäus, A. Kale, T. Chernenko, V. Torchilin and M. Diem, New ways of imaging uptake and intracellular fate of liposomal drug carrier systems inside individual cells, based on Raman microscopy, Molecular Pharmaceutics 5: (2) ((2008) ), 287–293, PMID: 18197626. |
36 | [[36]] M. Miljkovic, T. Chernenko, M.J. Romeo, B. Bird, C. Matthäus and M. Diem, Label-free imaging of human cells: Algorithms for image reconstruction of Raman hyperspectral datasets, Analyst 135: ((2010) ), 2002–2013. |
37 | [[37]] U. Neugebauer, T. Bocklitz, J.H. Clement, C. Krafft and J. Popp, Towards detection and identification of circulating tumour cells using Raman spectroscopy, Analyst 135: (12) ((2010) ), 3178–3182. |
38 | [[38]] U. Neugebauer, J.H. Clement, T. Bocklitz, C. Krafft and J. Popp, Identification and differentiation of single cells from peripheral blood by Raman spectroscopic imaging, Journal of Biophotonics 3: (8,9) ((2010) ), 579–587. |
39 | [[39]] K.C. Neuman, E.H. Chadd, G.F. Liou, K. Bergman and S.M. Block, Characterization of photodamage to Escherichia coli in optical traps, Biophysical Journal 77: (5) ((1999) ), 2856–2863. |
40 | [[40]] G.J. Puppels, J.H.F. Olminkhof, G.M.J. Segers-Nolten, C. Otto, F.F.M. De Mul and J. Greve, Laser irradiation and Raman spectroscopy of single living cells and chromosomes: Sample degradation occurs with 514.5 nm but not with 660 nm laser light, Experimental Cell Research 195: (2) ((1991) ), 361–367. |
41 | [[41]] R Core Team, R Development Core Team. R: A Language and Environment for Statistical Computing, R foundation for statistical computing, Vienna, Austria, (2014) . |
42 | [[42]] I.W. Schie, L. Alber, A.L. Gryshuk and J.W. Chan, Investigating drug induced changes in single, living lymphocytes based on Raman micro-spectroscopy, Analyst 139: (11) ((2014) ), 2726–2733. |
43 | [[43]] I.W. Schie and T. Huser, Methods and applications of Raman microspectroscopy to single-cell analysis, Appl. Spectrosc. 67: (8) ((2013) ), 813–828. |
44 | [[44]] U. Schmid, P. Rösch, M. Krause, M. Harz, J. Popp and K. Baumann, Gaussian mixture discriminant analysis for the single-cell differentiation of bacteria using micro-Raman spectroscopy, Chemometrics and Intelligent Laboratory Systems 96: (2) ((2009) ), 159–171. |
45 | [[45]] M. Schmitt and J. Popp, Raman spectroscopy at the beginning of the twenty-first century, J. Raman Spectrosc. 37: ((2006) ), 20–28. |
46 | [[46]] B. Schrader, Infrared and Raman Spectroscopy: Methods and Applications, Wiley, (2008) . |
47 | [[47]] N.M. Sijtsema, S.D. Wouters, C.J. De Grauw, C. Otto and J. Greve, Confocal direct imaging Raman microscope: Design and applications in biology, Appl. Spectrosc. 52: (3) ((1998) ), 348–355. |
48 | [[48]] E. Smith and G. Dent, Modern Raman Spectroscopy: A Practical Approach, Wiley, Hoboken, NJ, (2005) . |
49 | [[49]] R. Tabaksblat, R.J. Meier and B.J. Kip, Confocal Raman microspectroscopy: Theory and application to thin polymer samples, Appl. Spectrosc. 46: (1) ((1992) ), 60–68. |
50 | [[50]] A. Taleb, J. Diamond, J.J. McGarvey, J.R. Beattie, C. Toland and P.W. Hamilton, Raman microscopy for the chemometric analysis of tumor cells, The Journal of Physical Chemistry B 110: (39) ((2006) ), 19625–19631. |
51 | [[51]] J.M. Tedesco and K.L. Davis, Calibration of dispersive Raman process analyzers, in: Proceedings of SPIE, M. Fallahi, R.J. Nordstrom and T.R. Todd, eds, (1999) , pp. 200–212. |
52 | [[52]] H. Tsuda and J. Arends, Raman spectroscopy in dental research: A short review of recent studies, ADR 11: (4) ((1997) ), 539–547. |
53 | [[53]] N.A. Turko, I. Barnea, O. Blum, R. Korenstein and N.T. Shaked, Detection and controlled depletion of cancer cells using photothermal phase microscopy, Journal of Biophotonics 8: (9) ((2015) ), 755–763. |
54 | [[54]] N. Uzunbajakava and C. Otto, Combined Raman and continuous-wave-excited two-photon fluorescence cell imaging, Optics Letters 28: (21) ((2003) ), 2073–2075. |
55 | [[55]] K.P.J. Williams, G.D. Pitt, D.N. Batchelder and B.J. Kip, Confocal Raman microspectroscopy using a stigmatic spectrograph and CCD detector, Appl. Spectrosc. 48: (2) ((1994) ), 232–235. |
56 | [[56]] M.E. Winter, N-FINDR: An algorithm for fast autonomous spectral end-member determination in hyperspectral data, in: SPIE’s International Symposium on Optical Science, Engineering, and Instrumentation, International Society for Optics and Photonics, (1999) , pp. 266–275. |
57 | [[57]] C. Xie, J. Mace, M.A. Dinno, Y.Q. Li, W. Tang, R.J. Newton and P.J. Gemperline, Identification of single bacterial cells in aqueous solution using confocal laser tweezers Raman spectroscopy, Analytical Chemistry 77: (14) ((2005) ), 4390–4397. |




